| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-27 01:35:56 UTC |
|---|
| Update Date | 2016-11-09 01:22:30 UTC |
|---|
| Accession Number | CHEM041635 |
|---|
| Identification |
|---|
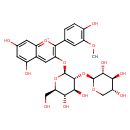
| Common Name | Peonidin 3-sambubioside |
|---|
| Class | Small Molecule |
|---|
| Description | A polyphenol metabolite detected in biological fluids |
|---|
| Contaminant Sources | - FooDB Chemicals
- HMDB Contaminants - Urine
|
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| Peonidin 3-O-sambubioside | HMDB |
|
|---|
| Chemical Formula | C27H31O15 |
|---|
| Average Molecular Mass | 595.526 g/mol |
|---|
| Monoisotopic Mass | 595.166 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | 3-{[(2S,3R,4S,5S,6R)-4,5-dihydroxy-6-(hydroxymethyl)-3-{[(2S,3R,4S,5R)-3,4,5-trihydroxyoxan-2-yl]oxy}oxan-2-yl]oxy}-5,7-dihydroxy-2-(4-hydroxy-3-methoxyphenyl)-1λ⁴-chromen-1-ylium |
|---|
| Traditional Name | 3-{[(2S,3R,4S,5S,6R)-4,5-dihydroxy-6-(hydroxymethyl)-3-{[(2S,3R,4S,5R)-3,4,5-trihydroxyoxan-2-yl]oxy}oxan-2-yl]oxy}-5,7-dihydroxy-2-(4-hydroxy-3-methoxyphenyl)-1λ⁴-chromen-1-ylium |
|---|
| SMILES | COC1=C(O)C=CC(=C1)C1=C(O[C@@H]2O[C@H](CO)[C@@H](O)[C@H](O)[C@H]2O[C@@H]2OC[C@@H](O)[C@H](O)[C@H]2O)C=C2C(O)=CC(O)=CC2=[O+]1 |
|---|
| InChI Identifier | InChI=1S/C27H30O15/c1-37-17-4-10(2-3-13(17)30)24-18(7-12-14(31)5-11(29)6-16(12)39-24)40-27-25(22(35)21(34)19(8-28)41-27)42-26-23(36)20(33)15(32)9-38-26/h2-7,15,19-23,25-28,32-36H,8-9H2,1H3,(H2-,29,30,31)/p+1/t15-,19-,20+,21-,22+,23-,25-,26+,27-/m1/s1 |
|---|
| InChI Key | PYUGATAPBPHGFK-UVSNCIQKSA-O |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as anthocyanidin-3-o-glycosides. These are phenolic compounds containing one anthocyanidin moiety which is O-glycosidically linked to a carbohydrate moiety at the C3-position. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Phenylpropanoids and polyketides |
|---|
| Class | Flavonoids |
|---|
| Sub Class | Flavonoid glycosides |
|---|
| Direct Parent | Anthocyanidin-3-O-glycosides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Anthocyanidin-3-o-glycoside
- Flavonoid-3-o-glycoside
- 3p-methoxyflavonoid-skeleton
- 4'-hydroxyflavonoid
- Hydroxyflavonoid
- 5-hydroxyflavonoid
- 7-hydroxyflavonoid
- Anthocyanidin
- Disaccharide
- Glycosyl compound
- O-glycosyl compound
- Benzopyran
- Methoxyphenol
- 1-benzopyran
- Anisole
- Phenoxy compound
- Methoxybenzene
- Phenol ether
- 1-hydroxy-4-unsubstituted benzenoid
- 1-hydroxy-2-unsubstituted benzenoid
- Alkyl aryl ether
- Phenol
- Oxane
- Benzenoid
- Monocyclic benzene moiety
- Heteroaromatic compound
- Secondary alcohol
- Acetal
- Organoheterocyclic compound
- Oxacycle
- Ether
- Polyol
- Organic oxygen compound
- Hydrocarbon derivative
- Alcohol
- Organooxygen compound
- Primary alcohol
- Organic cation
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00b9-5210290000-e3e9de62f248cebd6c56 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS ("Peonidin 3-sambubioside,1TMS,#1" TMS) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_7) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_8) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_9) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_7) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_8) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_9) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_10) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_11) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_12) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_13) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_14) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udj-0069520000-89b3e975bbc1f770b000 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-0149200000-371f563d33ed3d331f0d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0uk9-1987000000-d0569327e21d94bd1ad8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0i01-0003900000-69a386e53177c4575ee1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0udj-0209810000-426b51eb64e0c9a60227 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0002-2494110000-23de10cbfcff486abb47 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0041764 |
|---|
| FooDB ID | FDB029935 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | Not Available |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 101006858 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Neveu V, Perez-Jimenez J, Vos F, Crespy V, du Chaffaut L, Mennen L, Knox C, Eisner R, Cruz J, Wishart D, Scalbert A: Phenol-Explorer: an online comprehensive database on polyphenol contents in foods. Database (Oxford). 2010;2010:bap024. doi: 10.1093/database/bap024. Epub 2010 Jan 8. |
|
|---|