| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-27 01:19:07 UTC |
|---|
| Update Date | 2016-11-09 01:22:26 UTC |
|---|
| Accession Number | CHEM041268 |
|---|
| Identification |
|---|
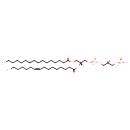
| Common Name | PGP(16:0/16:1(9Z)) |
|---|
| Class | Small Molecule |
|---|
| Description | PGP(16:0/16:1(9Z)) belongs to the class of glycerophosphoglycerophosphates, also called phosphatidylglycerophosphates (PGPs). These lipids contain a common glycerophosphate skeleton linked to at least one fatty acyl chain and a glycero-3-phosphate moiety. As is the case with diacylglycerols, phosphatidylglycerophosphates can have many different combinations of fatty acids of varying lengths and saturation attached to the C-1 and C-2 positions. PGP(16:0/16:1(9Z)), in particular, consists of one hexadecanoyl chain to the C-1 atom, and one 9Z-hexadecenoyl to the C-2 atom. In E. coli, PGPs can be found in the cytoplasmic membrane. The are synthesized by the addition of glycerol 3-phosphate to a CDP-diacylglycerol. In turn, PGPs are dephosphorylated to Phosphatidylglycerols (PGs) by the enzyme Phosphatidylglycerophosphatase. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 1-Palmitoyl-2-palmitoleoyl-sn-glycero-3-phospho-(1'-sn-glycerol-3'-phosphate) | HMDB | | 3-sn-Phosphatidyl-1'-sn-glycerol 3'-phosphoric acid | HMDB | | PGP(16:0/16:1) | HMDB | | PGP(16:0/16:1n7) | HMDB | | PGP(16:0/16:1W7) | HMDB | | PGP(32:1) | HMDB | | 1-Hexadecanoyl-2-(9Z-hexadecenoyl)-sn-glycero-3-phospho-(1'-sn-glycerol-3'-phosphate) | HMDB | | PGP(16:0/16:1(9Z)) | Lipid Annotator |
|
|---|
| Chemical Formula | C38H74O13P2 |
|---|
| Average Molecular Mass | 800.945 g/mol |
|---|
| Monoisotopic Mass | 800.460 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | [(2S)-3-({[(2R)-2-[(9Z)-hexadec-9-enoyloxy]-3-(hexadecanoyloxy)propoxy](hydroxy)phosphoryl}oxy)-2-hydroxypropoxy]phosphonic acid |
|---|
| Traditional Name | (2S)-3-{[(2R)-2-[(9Z)-hexadec-9-enoyloxy]-3-(hexadecanoyloxy)propoxy(hydroxy)phosphoryl]oxy}-2-hydroxypropoxyphosphonic acid |
|---|
| SMILES | [H][C@](O)(COP(O)(O)=O)COP(O)(=O)OC[C@@]([H])(COC(=O)CCCCCCCCCCCCCCC)OC(=O)CCCCCCC\C=C/CCCCCC |
|---|
| InChI Identifier | InChI=1S/C38H74O13P2/c1-3-5-7-9-11-13-15-17-19-21-23-25-27-29-37(40)47-33-36(34-50-53(45,46)49-32-35(39)31-48-52(42,43)44)51-38(41)30-28-26-24-22-20-18-16-14-12-10-8-6-4-2/h14,16,35-36,39H,3-13,15,17-34H2,1-2H3,(H,45,46)(H2,42,43,44)/b16-14-/t35-,36+/m0/s1 |
|---|
| InChI Key | RCAKEXBGCGLMOX-DCQLZOMZSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as phosphatidylglycerophosphates. These are glycerophosphoglycerophosphates in which two fatty acids are bonded to the 1-glycerol moiety through ester linkages. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Glycerophospholipids |
|---|
| Sub Class | Glycerophosphoglycerophosphates |
|---|
| Direct Parent | Phosphatidylglycerophosphates |
|---|
| Alternative Parents | |
|---|
| Substituents | - Diacylglycerophosphoglycerophosphate
- Sn-glycerol-3-phosphate
- Fatty acid ester
- Monoalkyl phosphate
- Dialkyl phosphate
- Dicarboxylic acid or derivatives
- Organic phosphoric acid derivative
- Phosphoric acid ester
- Alkyl phosphate
- Fatty acyl
- Secondary alcohol
- Carboxylic acid ester
- Carboxylic acid derivative
- Organooxygen compound
- Organic oxide
- Organic oxygen compound
- Alcohol
- Carbonyl group
- Hydrocarbon derivative
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0f6t-0490081530-55a138498138ecca372c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-053b-1591041200-e03a93cd0a01f3741cf8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-052k-4893221100-deb0aa426adf6174ca2d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-054t-3090030200-375cad12f6e125227779 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9050000000-a9baa34d7c44ba88a179 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9010000000-3d1708a487ebf6e3701b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0002-0010000900-1cf4f222fa3eff080c40 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0002-3091041400-b602219be812461b192d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0fba-3092061100-e4abaaab020b6afc20eb | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-2100062690-19d4b58b1fbdb56421e8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0k92-5600097400-2867fb8cdab51007291e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0k92-7496620000-c4d7ddfc0dce89035ba5 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0013473 |
|---|
| FooDB ID | FDB029473 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | Not Available |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 53481795 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | YMDB14374 |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|