| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-27 01:15:46 UTC |
|---|
| Update Date | 2016-11-09 01:22:25 UTC |
|---|
| Accession Number | CHEM041166 |
|---|
| Identification |
|---|
| Common Name | Nonanoylcarnitine |
|---|
| Class | Small Molecule |
|---|
| Description | A C9-acylcarnitine having nonanoyl as the acyl substituent. |
|---|
| Contaminant Sources | - FooDB Chemicals
- HMDB Contaminants - Urine
|
|---|
| Contaminant Type | Not Available |
|---|
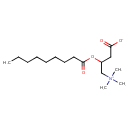
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 3-(Nonanoyloxy)-4-(trimethylammonio)butanoate | ChEBI | | 3-(Nonanoyloxy)-4-(trimethylammonio)butanoic acid | Generator | | Nonayl-L-carnitine | HMDB | | Nonaylcarnitine | HMDB |
|
|---|
| Chemical Formula | C16H31NO4 |
|---|
| Average Molecular Mass | 301.422 g/mol |
|---|
| Monoisotopic Mass | 301.225 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | 3-(nonanoyloxy)-4-(trimethylazaniumyl)butanoate |
|---|
| Traditional Name | 3-(nonanoyloxy)-4-(trimethylammonio)butanoate |
|---|
| SMILES | CCCCCCCCC(=O)OC(CC([O-])=O)C[N+](C)(C)C |
|---|
| InChI Identifier | InChI=1S/C16H31NO4/c1-5-6-7-8-9-10-11-16(20)21-14(12-15(18)19)13-17(2,3)4/h14H,5-13H2,1-4H3 |
|---|
| InChI Key | MPSPNFAQQQMFLK-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as acyl carnitines. These are organic compounds containing a fatty acid with the carboxylic acid attached to carnitine through an ester bond. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Fatty acid esters |
|---|
| Direct Parent | Acyl carnitines |
|---|
| Alternative Parents | |
|---|
| Substituents | - Acyl-carnitine
- Dicarboxylic acid or derivatives
- Tetraalkylammonium salt
- Quaternary ammonium salt
- Carboxylic acid ester
- Carboxylic acid salt
- Carboxylic acid derivative
- Carboxylic acid
- Organic nitrogen compound
- Organooxygen compound
- Organonitrogen compound
- Organic salt
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Organic oxygen compound
- Carbonyl group
- Amine
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00di-9200000000-5b0a4412e022a07b31a3 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0009000000-e69a13f756074c5f22a4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0f79-9005000000-9c86c8623895de0526be | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-9000000000-e9262cbaff8cb4ad0ba6 | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0013288 |
|---|
| FooDB ID | FDB029365 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 28533402 |
|---|
| ChEBI ID | 70997 |
|---|
| PubChem Compound ID | 53481699 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | YMDB01539 |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. https://www.ncbi.nlm.nih.gov/pubmed/?term=22770225 | | 2. Ferdinandusse S, Mulders J, IJlst L, Denis S, Dacremont G, Waterham HR, Wanders RJ: Molecular cloning and expression of human carnitine octanoyltransferase: evidence for its role in the peroxisomal beta-oxidation of branched-chain fatty acids. Biochem Biophys Res Commun. 1999 Sep 16;263(1):213-8. | | 3. Roux A, Xu Y, Heilier JF, Olivier MF, Ezan E, Tabet JC, Junot C: Annotation of the human adult urinary metabolome and metabolite identification using ultra high performance liquid chromatography coupled to a linear quadrupole ion trap-Orbitrap mass spectrometer. Anal Chem. 2012 Aug 7;84(15):6429-37. doi: 10.1021/ac300829f. Epub 2012 Jul 17. | | 4. Bahado-Singh RO, Ertl R, Mandal R, Bjorndahl TC, Syngelaki A, Han B, Dong E, Liu PB, Alpay-Savasan Z, Wishart DS, Nicolaides KH: Metabolomic prediction of fetal congenital heart defect in the first trimester. Am J Obstet Gynecol. 2014 Sep;211(3):240.e1-240.e14. doi: 10.1016/j.ajog.2014.03.056. Epub 2014 Apr 1. | | 5. Simons K, Toomre D: Lipid rafts and signal transduction. Nat Rev Mol Cell Biol. 2000 Oct;1(1):31-9. | | 6. Watson AD: Thematic review series: systems biology approaches to metabolic and cardiovascular disorders. Lipidomics: a global approach to lipid analysis in biological systems. J Lipid Res. 2006 Oct;47(10):2101-11. Epub 2006 Aug 10. | | 7. Sethi JK, Vidal-Puig AJ: Thematic review series: adipocyte biology. Adipose tissue function and plasticity orchestrate nutritional adaptation. J Lipid Res. 2007 Jun;48(6):1253-62. Epub 2007 Mar 20. | | 8. Lingwood D, Simons K: Lipid rafts as a membrane-organizing principle. Science. 2010 Jan 1;327(5961):46-50. doi: 10.1126/science.1174621. | | 9. Cho K, Moon JS, Kang JH, Jang HB, Lee HJ, Park SI, Yu KS, Cho JY: Combined untargeted and targeted metabolomic profiling reveals urinary biomarkers for discriminating obese from normal-weight adolescents. Pediatr Obes. 2017 Apr;12(2):93-101. doi: 10.1111/ijpo.12114. Epub 2016 Feb 22. | | 10. FRITZ IB: Action of carnitine on long chain fatty acid oxidation by liver. Am J Physiol. 1959 Aug;197:297-304. doi: 10.1152/ajplegacy.1959.197.2.297. | | 11. Violante S, Achetib N, van Roermund CWT, Hagen J, Dodatko T, Vaz FM, Waterham HR, Chen H, Baes M, Yu C, Argmann CA, Houten SM: Peroxisomes can oxidize medium- and long-chain fatty acids through a pathway involving ABCD3 and HSD17B4. FASEB J. 2019 Mar;33(3):4355-4364. doi: 10.1096/fj.201801498R. Epub 2018 Dec 12. | | 12. Chen C, Hou G, Zeng C, Ren Y, Chen X, Peng C: Metabolomic profiling reveals amino acid and carnitine alterations as metabolic signatures in psoriasis. Theranostics. 2021 Jan 1;11(2):754-767. doi: 10.7150/thno.51154. eCollection 2021. | | 13. Ottas A, Fishman D, Okas TL, Kingo K, Soomets U: The metabolic analysis of psoriasis identifies the associated metabolites while providing computational models for the monitoring of the disease. Arch Dermatol Res. 2017 Sep;309(7):519-528. doi: 10.1007/s00403-017-1760-1. Epub 2017 Jul 10. | | 14. The lipid handbook with CD-ROM |
|
|---|