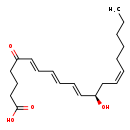
5-Oxo-6-trans-leukotriene B4 (CHEM041001)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2016-05-27 01:07:44 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2016-11-09 01:22:22 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Accession Number | CHEM041001 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | 5-Oxo-6-trans-leukotriene B4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Small Molecule | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | 5-oxo-6-trans-leukotriene B4 is the metabolite of lipid omega-oxidation of leukotriene B4 (LTB4). LTB4 is the major metabolite in neutrophil polymorphonuclear leukocytes. Omega-oxidation is the major pathway for the catabolism of leukotriene B4 in human polymorphonuclear leukocytes. Leukotrienes are metabolites of arachidonic acid derived from the action of 5-LO (5-lipoxygenase). The immediate product of 5-LO is LTA4 (leukotriene A4), which is enzymatically converted into either LTB4 (leukotriene B4) by LTA4 hydrolase or LTC4 (leukotriene C4) by LTC4 synthase. The regulation of leukotriene production occurs at various levels, including expression of 5-LO, translocation of 5-LO to the perinuclear region, and phosphorylation to either enhance or inhibit the activity of 5-LO. Biologically active LTB4 is metabolized by omega-oxidation carried out by specific cytochrome P450s (CYP4F) followed by beta-oxidation from the omega-carboxy position and after CoA ester formation (PMID: 7649996, 17623009, 2853166, 6088485). Leukotrienes are eicosanoids. The eicosanoids consist of the prostaglandins (PGs), thromboxanes (TXs), leukotrienes (LTs), and lipoxins (LXs). The PGs and TXs are collectively identified as prostanoids. Prostaglandins were originally shown to be synthesized in the prostate gland, thromboxanes from platelets (thrombocytes), and leukotrienes from leukocytes, hence the derivation of their names. All mammalian cells except erythrocytes synthesize eicosanoids. These molecules are extremely potent, able to cause profound physiological effects at very dilute concentrations. All eicosanoids function locally at the site of synthesis, through receptor-mediated G-protein linked signalling pathways. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Contaminant Sources |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Contaminant Type | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C20H30O4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Mass | 334.450 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Mass | 334.214 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | (6E,8E,10E,12R,14Z)-12-hydroxy-5-oxoicosa-6,8,10,14-tetraenoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | (6E,8E,10E,12R,14Z)-12-hydroxy-5-oxoicosa-6,8,10,14-tetraenoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | CCCCC\C=C/C[C@@H](O)\C=C\C=C\C=C\C(=O)CCCC(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1S/C20H30O4/c1-2-3-4-5-6-9-13-18(21)14-10-7-8-11-15-19(22)16-12-17-20(23)24/h6-11,14-15,18,21H,2-5,12-13,16-17H2,1H3,(H,23,24)/b8-7+,9-6-,14-10+,15-11+/t18-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | MLZJFLKEKVDNAZ-BEWISGCMSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as leukotrienes. These are eicosanoids containing a hydroxyl group attached to the aliphatic chain of an arachidonic acid. Leukotrienes have four double bonds, three (and only three) of which are conjugated. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Lipids and lipid-like molecules | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Fatty Acyls | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Eicosanoids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Leukotrienes | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Status | Detected and Not Quantified | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Origin | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biofluid Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tissue Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Applications | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Roles | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Roles | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Appearance | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Profile | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Route of Exposure | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mechanism of Toxicity | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolism | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Values | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lethal Dose | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Carcinogenicity (IARC Classification) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Uses/Sources | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Minimum Risk Level | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Health Effects | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symptoms | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Treatment | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | HMDB0012824 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FooDB ID | FDB029181 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Phenol Explorer ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KNApSAcK ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BiGG ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| METLIN ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemspider ID | 30776646 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound ID | 53481526 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kegg Compound ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| YMDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ECMDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MSDS | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||