| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-27 01:07:07 UTC |
|---|
| Update Date | 2016-11-09 01:22:22 UTC |
|---|
| Accession Number | CHEM040981 |
|---|
| Identification |
|---|
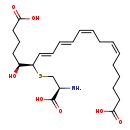
| Common Name | 20-COOH-leukotriene E4 |
|---|
| Class | Small Molecule |
|---|
| Description | 20-COOH-leukotriene E4 is a metabolite through lipid oxidation of Leukotriene E4 (LTE4).Leukotriene E4 (LTE4) is a cysteinyl leukotriene. Cysteinyl leukotrienes (CysLTs) are a family of potent inflammatory mediators that appear to contribute to the pathophysiologic features of allergic rhinitis. Nasal blockage induced by CysLTs is mainly due to dilatation of nasal blood vessels, which can be induced by the nitric oxide produced through CysLT1 receptor activation. LTE4, activate contractile and inflammatory processes via specific interaction with putative seven transmembrane-spanning receptors that couple to G proteins and subsequent intracellular signaling pathways. LTE4 is metabolized from leukotriene C4 in a reaction catalyzed by gamma-glutamyl transpeptidase and a particulate dipeptidase from kidney. (PMID: 12607939, 12432945, 6311078). Leukotrienes are eicosanoids. The eicosanoids consist of the prostaglandins (PGs), thromboxanes (TXs), leukotrienes (LTs), and lipoxins (LXs). The PGs and TXs are collectively identified as prostanoids. Prostaglandins were originally shown to be synthesized in the prostate gland, thromboxanes from platelets (thrombocytes), and leukotrienes from leukocytes, hence the derivation of their names. All mammalian cells except erythrocytes synthesize eicosanoids. These molecules are extremely potent, able to cause profound physiological effects at very dilute concentrations. All eicosanoids function locally at the site of synthesis, through receptor-mediated G-protein linked signalling pathways. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 20-COOH-LTE4 | HMDB | | 6-(S-Cysteinyl)-(5S)-hydroxy-(7E,9E,11Z,14Z)-eicosatetraenedioate | HMDB | | 6-(S-Cysteinyl)-(5S)-hydroxy-(7E,9E,11Z,14Z)-eicosatetraenedioic acid | HMDB |
|
|---|
| Chemical Formula | C23H35NO7S |
|---|
| Average Molecular Mass | 469.592 g/mol |
|---|
| Monoisotopic Mass | 469.213 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | (6Z,9Z,11E,13E,15R,16S)-15-{[(2S)-2-amino-2-carboxyethyl]sulfanyl}-16-hydroxyicosa-6,9,11,13-tetraenedioic acid |
|---|
| Traditional Name | (6Z,9Z,11E,13E,15R,16S)-15-{[(2S)-2-amino-2-carboxyethyl]sulfanyl}-16-hydroxyicosa-6,9,11,13-tetraenedioic acid |
|---|
| SMILES | N[C@H](CS[C@H](\C=C\C=C\C=C/C\C=C/CCCCC(O)=O)[C@@H](O)CCCC(O)=O)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C23H35NO7S/c24-18(23(30)31)17-32-20(19(25)13-12-16-22(28)29)14-10-8-6-4-2-1-3-5-7-9-11-15-21(26)27/h2-6,8,10,14,18-20,25H,1,7,9,11-13,15-17,24H2,(H,26,27)(H,28,29)(H,30,31)/b4-2-,5-3-,8-6+,14-10+/t18-,19+,20-/m1/s1 |
|---|
| InChI Key | HVLRBLGTGJGVCX-RHSCJZQUSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as cysteine and derivatives. Cysteine and derivatives are compounds containing cysteine or a derivative thereof resulting from reaction of cysteine at the amino group or the carboxy group, or from the replacement of any hydrogen of glycine by a heteroatom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Cysteine and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Cysteine or derivatives
- S-alkyl-l-cysteine
- Alpha-amino acid
- D-alpha-amino acid
- Tricarboxylic acid or derivatives
- Hydroxy fatty acid
- Thia fatty acid
- Fatty acyl
- Secondary alcohol
- Amino acid
- Carboxylic acid
- Dialkylthioether
- Sulfenyl compound
- Thioether
- Alcohol
- Hydrocarbon derivative
- Organic oxide
- Primary amine
- Organosulfur compound
- Organooxygen compound
- Organonitrogen compound
- Primary aliphatic amine
- Organopnictogen compound
- Organic oxygen compound
- Carbonyl group
- Amine
- Organic nitrogen compound
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0ukc-3219600000-afc81424c577c7c73830 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (3 TMS) - 70eV, Positive | splash10-00dl-8400918000-987eca40d2a4c9ab31e8 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0ue9-0001900000-a9aa6251a352d0e8e2a6 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0lk9-1249700000-127f1da4ed2b24fdf00f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0cmv-4019200000-c4f5f9fd8da1131bd7f4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0gir-0103900000-46c830f394a499c0440d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-03l9-1219200000-0336064a6d9bcaf9f0c9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-000i-9302000000-10f31ff39e1d782b7a27 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0ul0-0029500000-a97dae5c18a6435b7889 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0kur-0039500000-4b7053be1be20c285946 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-01e9-4910000000-2ad4c146132a94fd0587 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-01x0-2309400000-bcb027d5a55752e9a430 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-03di-1219000000-43c90b9b27b55393b866 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-03xr-5559000000-5e7c46a6b1b8554d75a6 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0012634 |
|---|
| FooDB ID | FDB029159 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 30776643 |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 53481508 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Mizutani N: [Studies on the experimental allergic rhinitis induced by Japanese cedar pollen--role of cysteinyl leukotrienes in nasal allergic symptoms]. Yakugaku Zasshi. 2003 Jan;123(1):1-8. | | 2. Evans JF: Cysteinyl leukotriene receptors. Prostaglandins Other Lipid Mediat. 2002 Aug;68-69:587-97. | | 3. Hammarstrom S: Leukotrienes. Annu Rev Biochem. 1983;52:355-77. | | 4. Brunk E, Sahoo S, Zielinski DC, Altunkaya A, Drager A, Mih N, Gatto F, Nilsson A, Preciat Gonzalez GA, Aurich MK, Prlic A, Sastry A, Danielsdottir AD, Heinken A, Noronha A, Rose PW, Burley SK, Fleming RMT, Nielsen J, Thiele I, Palsson BO: Recon3D enables a three-dimensional view of gene variation in human metabolism. Nat Biotechnol. 2018 Mar;36(3):272-281. doi: 10.1038/nbt.4072. Epub 2018 Feb 19. |
|
|---|