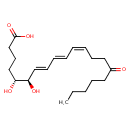
13,14-Dihydro-15-oxo-lipoxin A4 (CHEM040953)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2016-05-27 01:05:34 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2016-11-09 01:22:21 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Accession Number | CHEM040953 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | 13,14-Dihydro-15-oxo-lipoxin A4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Small Molecule | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | 13,14-dihydro-15-oxo-lipoxin A4 is a lipoxin derivative. Lipoxins (LXs) and aspirin-triggered Lipoxin (ATL) are trihydroxytetraene-containing eicosanoids generated from arachidonic acid that are distinct in structure, formation, and function from the many other proinflammatory lipid-derived mediators. These endogenous eicosanoids have now emerged as founding members of the first class of lipid/chemical mediators involved in the resolution of the inflammatory response. Lipoxin A4 (LXA4), ATL, and their metabolic stable analogs elicit cellular responses and regulate leukocyte trafficking in vivo by activating the specific receptor, ALX. Many of the eicosanoids derived from arachidonic acid (AA2), including prostaglandins (PGs) and leukotrienes (LTs), play important roles as local mediators exerting a wide range of actions relevant in immune hypersensitivity and inflammation. However, recent observations indicate that other agents derived from the lipoxygenase (LO) pathways are formed and play a key role in initiating the resolution of acute inflammation. This phenomenon is an active process that is governed by specific lipid mediators and involves a series of well-orchestrated temporal events. Thus, potent locally released mediators serve as checkpoint controllers of inflammation. In addition to the well-appreciated ability of aspirin to inhibit PGs, aspirin also acetylates cyclooxygenase (COX)-2, triggering the formation of a 15-epimeric form of lipoxins, termed aspirin-triggered LXA4 (ATL). These eicosanoids (i.e., LXA4 and ATL) with a unique trihydroxytetraene structure function as 'stop signals' in inflammation and actively participate in dampening host responses to bring the inflammation to a close, namely, resolution. LXA4 and ATL elicit the multicellular responses via a specific G protein-coupled receptor (GPCR) termed ALX that has been identified in human. (PMID: 16968948, 11478982). | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Contaminant Sources |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Contaminant Type | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C20H32O5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Mass | 352.465 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Mass | 352.225 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | (5R,6R,7E,9E,11Z)-5,6-dihydroxy-15-oxoicosa-7,9,11-trienoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | (5R,6R,7E,9E,11Z)-5,6-dihydroxy-15-oxoicosa-7,9,11-trienoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | CCCCCC(=O)CC\C=C/C=C/C=C/[C@@H](O)[C@H](O)CCCC(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1S/C20H32O5/c1-2-3-8-12-17(21)13-9-6-4-5-7-10-14-18(22)19(23)15-11-16-20(24)25/h4-7,10,14,18-19,22-23H,2-3,8-9,11-13,15-16H2,1H3,(H,24,25)/b6-4-,7-5+,14-10+/t18-,19-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | FPRPRBFSKMFXRV-HJGGDGFVSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as hydroxyeicosatrienoic acids. These are eicosanoic acids with an attached hydroxyl group and three CC double bonds. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Lipids and lipid-like molecules | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Fatty Acyls | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Eicosanoids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Hydroxyeicosatrienoic acids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Status | Detected and Not Quantified | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Origin | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biofluid Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tissue Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Applications | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Roles | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Roles | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Appearance | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Profile | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Route of Exposure | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mechanism of Toxicity | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolism | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Values | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lethal Dose | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Carcinogenicity (IARC Classification) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Uses/Sources | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Minimum Risk Level | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Health Effects | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symptoms | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Treatment | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | HMDB0012564 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FooDB ID | FDB029130 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Phenol Explorer ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KNApSAcK ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BiGG ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| METLIN ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemspider ID | 30776637 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound ID | 53481470 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kegg Compound ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| YMDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ECMDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MSDS | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||