| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-27 00:30:32 UTC |
|---|
| Update Date | 2016-11-09 01:22:19 UTC |
|---|
| Accession Number | CHEM040745 |
|---|
| Identification |
|---|
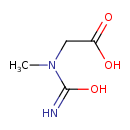
| Common Name | N-Carbamoylsarcosine |
|---|
| Class | Small Molecule |
|---|
| Description | The N-carbamoyl derivative of sarcosine. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| Carbamoyl sarcosine | HMDB | | Carbamoyl-sarcosine | HMDB | | CMS | HMDB | | [Carbamoyl(methyl)amino]acetic acid | HMDB |
|
|---|
| Chemical Formula | C4H8N2O3 |
|---|
| Average Molecular Mass | 132.118 g/mol |
|---|
| Monoisotopic Mass | 132.053 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | 2-[(C-hydroxycarbonimidoyl)(methyl)amino]acetic acid |
|---|
| Traditional Name | [C-hydroxycarbonimidoyl(methyl)amino]acetic acid |
|---|
| SMILES | CN(CC(O)=O)C(O)=N |
|---|
| InChI Identifier | InChI=1S/C4H8N2O3/c1-6(4(5)9)2-3(7)8/h2H2,1H3,(H2,5,9)(H,7,8) |
|---|
| InChI Key | SREKYKXYSQMOIB-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as alpha amino acids and derivatives. These are amino acids in which the amino group is attached to the carbon atom immediately adjacent to the carboxylate group (alpha carbon), or a derivative thereof. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Alpha amino acids and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Alpha-amino acid or derivatives
- Urea
- Carbonic acid derivative
- Monocarboxylic acid or derivatives
- Carboxylic acid
- Organic nitrogen compound
- Organic oxygen compound
- Organopnictogen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Organonitrogen compound
- Carbonyl group
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-059l-9420000000-26d7547dadb70ab4fc08 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0006-9100000000-f7eef7f30a71b2437a21 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-4900000000-828e01495f55c3ba6112 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-006x-9100000000-d5aaf1e2016e0db5e1ef | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00di-9000000000-7b8ae0e68a69def64830 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001c-9300000000-64fe3175ab61247e7c9c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000l-9100000000-3ef98ae24f347887d27a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9000000000-116ba20e6e0fcf01e7db | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-9000000000-df2ca252dc264c27f09e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000i-9000000000-87f7d4792f3b20ebdd54 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-006x-9000000000-1e503e67b415a9fae1b0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000f-9100000000-20d310b987c7d3c62e72 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0076-9000000000-7debac7e5b881e478ecc | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0006-9000000000-a06e49ac0ea6780c12d1 | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0012265 |
|---|
| FooDB ID | FDB028903 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | CARBAMOYL-SARCOSINE |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 388495 |
|---|
| ChEBI ID | 15737 |
|---|
| PubChem Compound ID | 439375 |
|---|
| Kegg Compound ID | C01043 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | ECMDB23800 |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Jiang S, Sun X, Zhang S: The ycaC-related protein from the amphioxus Branchiostoma belcheri (BbycaCR) interacts with creatine kinase. FEBS J. 2008 Sep;275(18):4597-605. doi: 10.1111/j.1742-4658.2008.06602.x. Epub 2008 Aug 11. | | 2. Wang WC, Hsu WH, Chien FT, Chen CY: Crystal structure and site-directed mutagenesis studies of N-carbamoyl-D-amino-acid amidohydrolase from Agrobacterium radiobacter reveals a homotetramer and insight into a catalytic cleft. J Mol Biol. 2001 Feb 16;306(2):251-61. | | 3. Lemaitre N, Callebaut I, Frenois F, Jarlier V, Sougakoff W: Study of the structure-activity relationships for the pyrazinamidase (PncA) from Mycobacterium tuberculosis. Biochem J. 2001 Feb 1;353(Pt 3):453-8. | | 4. Kim JM, Shimizu S, Yamada H: Purification and characterization of a novel enzyme, N-carbamoylsarcosine amidohydrolase, from Pseudomonas putida 77. J Biol Chem. 1986 Sep 5;261(25):11832-9. | | 5. Parsons JF, Calabrese K, Eisenstein E, Ladner JE: Structure and mechanism of Pseudomonas aeruginosa PhzD, an isochorismatase from the phenazine biosynthetic pathway. Biochemistry. 2003 May 20;42(19):5684-93. | | 6. Zajc A, Romao MJ, Turk B, Huber R: Crystallographic and fluorescence studies of ligand binding to N-carbamoylsarcosine amidohydrolase from Arthrobacter sp. J Mol Biol. 1996 Oct 25;263(2):269-83. | | 7. Kim JM, Shimizu S, Yamada H: Amidohydrolysis of N-methylhydantoin coupled with ATP hydrolysis. Biochem Biophys Res Commun. 1987 Feb 13;142(3):1006-12. | | 8. Romao MJ, Turk D, Gomis-Ruth FX, Huber R, Schumacher G, Mollering H, Russmann L: Crystal structure analysis, refinement and enzymatic reaction mechanism of N-carbamoylsarcosine amidohydrolase from Arthrobacter sp. at 2.0 A resolution. J Mol Biol. 1992 Aug 20;226(4):1111-30. | | 9. Ogawa J, Nirdnoy W, Tabata M, Yamada H, Shimizu S: A new enzymatic method for the measurement of creatinine involving a novel ATP-dependent enzyme, N-methylhydantoin amidohydrolase. Biosci Biotechnol Biochem. 1995 Dec;59(12):2292-4. | | 10. Hermann M, Knerr HJ, Mai N, Gross A, Kaltwasser H: Creatinine and N-methylhydantoin degradation in two newly isolated Clostridium species. Arch Microbiol. 1992;157(5):395-401. | | 11. Luo HB, Zheng H, Zimmerman MD, Chruszcz M, Skarina T, Egorova O, Savchenko A, Edwards AM, Minor W: Crystal structure and molecular modeling study of N-carbamoylsarcosine amidase Ta0454 from Thermoplasma acidophilum. J Struct Biol. 2010 Mar;169(3):304-11. doi: 10.1016/j.jsb.2009.11.008. Epub 2009 Nov 20. | | 12. Ogawa J, Shimizu S, Yamada H: N-carbamoyl-D-amino acid amidohydrolase from Comamonas sp. E222c purification and characterization. Eur J Biochem. 1993 Mar 15;212(3):685-91. | | 13. Caruthers J, Zucker F, Worthey E, Myler PJ, Buckner F, Van Voorhuis W, Mehlin C, Boni E, Feist T, Luft J, Gulde S, Lauricella A, Kaluzhniy O, Anderson L, Le Trong I, Holmes MA, Earnest T, Soltis M, Hodgson KO, Hol WG, Merritt EA: Crystal structures and proposed structural/functional classification of three protozoan proteins from the isochorismatase superfamily. Protein Sci. 2005 Nov;14(11):2887-94. Epub 2005 Sep 30. | | 14. Shimizu S, Kim JM, Yamada H: Microbial enzymes for creatinine assay: a review. Clin Chim Acta. 1989 Dec 15;185(3):241-52. |
|
|---|