| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 07:18:40 UTC |
|---|
| Update Date | 2016-11-09 01:21:27 UTC |
|---|
| Accession Number | CHEM036197 |
|---|
| Identification |
|---|
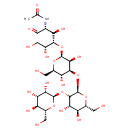
| Common Name | alpha-D-Manp-(1->2)-alpha-D-Manp-(1->2)-alpha-D-Manp-(1->3)-beta-D-Manp-(1->4)-D-GlcNAcp |
|---|
| Class | Small Molecule |
|---|
| Description | Alpha-D-Manp-(1 -> 2)-a-D-Manp-(1 -> 2)-a-D-Manp-(1 -> 3)-b-D-Manp-(1 -> 4)-D-GlcNAcp is a pentasaccharide found in the urine of patients with mannosidosis (PMID: 953057). It is a glycan structure of human milk glycoprotein glycopeptides (PMID: 6698017). This oligosaccharide is derived from high-mannose and different complex and hybrid asparagine-linked glycans and from the storage products in alpha-mannosidosis. The human liver lysosomal alpha-mannosidase (EC 3.2.1.24) enzyme that specifically hydrolyzes all alpha(1-2)-, alpha(1-3)- and alpha(1-6)-mannosidic linkages in these glycans. (PMID: 1872811). |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| a-D-Manp-(1 -> 2)-a-D-manp-(1 -> 2)-a-D-manp-(1 -> 3)-b-D-manp-(1 -> 4)-D-glcnacp | Generator | | Α-D-manp-(1 -> 2)-a-D-manp-(1 -> 2)-a-D-manp-(1 -> 3)-b-D-manp-(1 -> 4)-D-glcnacp | Generator | | a-D-Manp-(1 -> 2)-a-D-manp-(1 -> 2)-a-D-manp-(1 -> 3)-b-D-manp-(1 -> 4)-D-glcnacpo-alpha-D-mannopyranosyl-(1->2)-O-alpha-D-mannopyranosyl-(1->3)-O-beta-D-mannopyranosyl-(1->4)-2-(acetylamino)-2-deoxy- D-glucose | HMDB | | alpha-D-Manp-(1 -> 2)-alpha-D-manp-(1 -> 2)-alpha-D-manp-(1 -> 3)-beta-D-manp-(1 -> 4)-D-glcnacpo-alpha-D-mannopyranosyl-(1->2)-O-alpha-D-mannopyranosyl-(1->3)-O-beta-D-mannopyranosyl-(1->4)-2-(acetylamino)-2-deoxy- D-glucose | HMDB | | O-alpha-delta-Mannopyranosyl-(1->2)-O-alpha-delta-mannopyranosyl-(1->3)-O-beta-delta-mannopyranosyl-(1->4)-2-(acetylamino)-2-deoxy- D-glucose | HMDB | | N-[(2R,3R,4S,5R)-4-{[(2S,3S,4S,5R,6R)-4-{[(2R,4S,5S,6R)-4,5-dihydroxy-6-(hydroxymethyl)-3-{[(2R,3S,4S,5S,6R)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}oxan-2-yl]oxy}-3,5-dihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}-3,5,6-trihydroxy-1-oxohexan-2-yl]ethanimidate | Generator, HMDB | | a-D-Manp-(1->2)-a-D-manp-(1->2)-a-D-manp-(1->3)-b-D-manp-(1->4)-D-glcnacp | Generator | | Α-D-manp-(1->2)-α-D-manp-(1->2)-α-D-manp-(1->3)-β-D-manp-(1->4)-D-glcnacp | Generator |
|
|---|
| Chemical Formula | C26H45NO21 |
|---|
| Average Molecular Mass | 707.630 g/mol |
|---|
| Monoisotopic Mass | 707.248 g/mol |
|---|
| CAS Registry Number | 52134-33-5 |
|---|
| IUPAC Name | N-[(2R,3R,4S,5R)-4-{[(2S,3S,4S,5R,6R)-4-{[(2R,4S,5S,6R)-4,5-dihydroxy-6-(hydroxymethyl)-3-{[(2R,3S,4S,5S,6R)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}oxan-2-yl]oxy}-3,5-dihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}-3,5,6-trihydroxy-1-oxohexan-2-yl]acetamide |
|---|
| Traditional Name | N-[(2R,3R,4S,5R)-4-{[(2S,3S,4S,5R,6R)-4-{[(2R,4S,5S,6R)-4,5-dihydroxy-6-(hydroxymethyl)-3-{[(2R,3S,4S,5S,6R)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}oxan-2-yl]oxy}-3,5-dihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}-3,5,6-trihydroxy-1-oxohexan-2-yl]acetamide |
|---|
| SMILES | CC(=O)N[C@@H](C=O)[C@@H](O)[C@H](O[C@@H]1O[C@H](CO)[C@@H](O)[C@H](O[C@H]2O[C@H](CO)[C@@H](O)[C@H](O)C2O[C@H]2O[C@H](CO)[C@@H](O)[C@H](O)[C@@H]2O)[C@@H]1O)[C@H](O)CO |
|---|
| InChI Identifier | InChI=1S/C26H45NO21/c1-7(33)27-8(2-28)13(35)21(9(34)3-29)46-25-20(42)22(16(38)12(6-32)44-25)47-26-23(18(40)15(37)11(5-31)45-26)48-24-19(41)17(39)14(36)10(4-30)43-24/h2,8-26,29-32,34-42H,3-6H2,1H3,(H,27,33)/t8-,9+,10+,11+,12+,13+,14+,15+,16+,17-,18-,19-,20-,21+,22-,23?,24+,25-,26+/m0/s1 |
|---|
| InChI Key | POROYQINIQHUGJ-APJJTYCFSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as oligosaccharides. These are carbohydrates made up of 3 to 10 monosaccharide units linked to each other through glycosidic bonds. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Oligosaccharides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Oligosaccharide
- Fatty acyl glycoside
- Alkyl glycoside
- Glycosyl compound
- O-glycosyl compound
- Amino saccharide
- Beta-hydroxy aldehyde
- Oxane
- Fatty acyl
- Acetamide
- Secondary carboxylic acid amide
- Secondary alcohol
- Carboxamide group
- Organoheterocyclic compound
- Polyol
- Acetal
- Oxacycle
- Carboxylic acid derivative
- Aldehyde
- Organopnictogen compound
- Organic oxide
- Hydrocarbon derivative
- Carbonyl group
- Organonitrogen compound
- Organic nitrogen compound
- Alcohol
- Primary alcohol
- Aliphatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aliphatic heteromonocyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-03fr-4201009000-c6e66f8c14f635ec35a8 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_7) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_8) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_9) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_10) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_11) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_12) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_13) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_14) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_15) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_7) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_8) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_9) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0g7m-1298076100-fbbcf6d368dbfd23d88a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-5965031000-6eca7dcd817dc7bfa4b4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0h90-5895112000-65f1f72d04c1493d4e9f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004r-2411049100-8e3078875dc766a955ef | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0kdr-4932126000-4f2ee786d727032ffc94 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-00b9-5942000000-cf4dc592b49043b4ac96 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a5i-1100029200-3dd477a70746e5e07478 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a59-9000018000-c3821bde3976d5552037 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-052f-9200011000-722711dbd6bb1da308c3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0abm-0210019200-20c2d8c3a16452738a07 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0c0s-9800016200-9f35a70b731460efd89e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a5l-9200001000-c9aee2c23d79b119f4f3 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0006699 |
|---|
| FooDB ID | FDB024031 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 35016016 |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 53477881 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. al Daher S, de Gasperi R, Daniel P, Hall N, Warren CD, Winchester B: The substrate-specificity of human lysosomal alpha-D-mannosidase in relation to genetic alpha-mannosidosis. Biochem J. 1991 Aug 1;277 ( Pt 3):743-51. | | 2. Yanagida K, Natsuka S, Hase S: Structural diversity of cytosolic free oligosaccharides in the human hepatoma cell line, HepG2. Glycobiology. 2006 Apr;16(4):294-304. Epub 2005 Dec 27. | | 3. De Gasperi R, Daniel PF, Warren CD: A human lysosomal alpha-mannosidase specific for the core of complex glycans. J Biol Chem. 1992 May 15;267(14):9706-12. | | 4. Strecker G, Fournet B, Bouquelet S, Montreuil J, Dhondt JL, Farriaux JP: [Chemistry of urinary mannosides excreted in mannosidosis]. Biochimie. 1976;58(5):579-86. | | 5. Pierce-Cretel A, Debray H, Montreuil J, Spik G, Van Halbeek H, Mutsaers JH, Vliegenthart JF: Primary structure of N-glycosidically linked asialoglycans of secretory immunoglobulins A from human milk. Eur J Biochem. 1984 Mar 1;139(2):337-49. |
|
|---|