| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 07:11:16 UTC |
|---|
| Update Date | 2016-11-09 01:21:25 UTC |
|---|
| Accession Number | CHEM036052 |
|---|
| Identification |
|---|
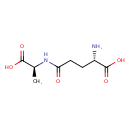
| Common Name | 5-L-Glutamyl-L-alanine |
|---|
| Class | Small Molecule |
|---|
| Description | A gamma-glutamylalanine obtained by formal condensation of the gamma-carboxy group of L-glutamic acid with the amino group of L-alanine. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 5-L-Glutamyl-L-alanine | ChEBI | | gamma-L-Glutamyl-L-alanine | ChEBI | | L-gamma-Glu-L-ala | ChEBI | | N-L-gamma-Glutamyl-L-alanine | ChEBI | | g-L-Glutamyl-L-alanine | Generator | | Γ-L-glutamyl-L-alanine | Generator | | L-g-Glu-L-ala | Generator | | L-Γ-glu-L-ala | Generator | | N-L-g-Glutamyl-L-alanine | Generator | | N-L-Γ-glutamyl-L-alanine | Generator | | g-Glutamylalanine | Generator | | Γ-glutamylalanine | Generator | | γ-Glu-Ala | HMDB | | γ-L-Glu-L-Ala | HMDB | | L-γ-Glutamyl-L-alanine | HMDB | | N-γ-Glutamylalanine | HMDB | | N-L-γ-Glutamylalanine | HMDB | | gamma-Glu-Ala | HMDB | | gamma-L-Glu-L-Ala | HMDB | | L-gamma-Glutamyl-L-alanine | HMDB | | N-gamma-Glutamylalanine | HMDB | | N-L-gamma-Glutamylalanine | HMDB | | gamma-Glutamylalanine | HMDB, ChEBI | | 5-L-Glutamylalanine | HMDB | | g-Glu-Ala | HMDB | | N-γ-L-Glutamyl-L-alanine | HMDB | | N-gamma-L-Glutamyl-L-alanine | HMDB |
|
|---|
| Chemical Formula | C8H14N2O5 |
|---|
| Average Molecular Mass | 218.207 g/mol |
|---|
| Monoisotopic Mass | 218.090 g/mol |
|---|
| CAS Registry Number | 5875-41-2 |
|---|
| IUPAC Name | (2S)-2-amino-4-{[(1S)-1-carboxyethyl]carbamoyl}butanoic acid |
|---|
| Traditional Name | g-glu-ala |
|---|
| SMILES | C[C@H](NC(=O)CC[C@H](N)C(O)=O)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C8H14N2O5/c1-4(7(12)13)10-6(11)3-2-5(9)8(14)15/h4-5H,2-3,9H2,1H3,(H,10,11)(H,12,13)(H,14,15)/t4-,5-/m0/s1 |
|---|
| InChI Key | WQXXXVRAFAKQJM-WHFBIAKZSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as dipeptides. These are organic compounds containing a sequence of exactly two alpha-amino acids joined by a peptide bond. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Dipeptides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Alpha-dipeptide
- Gamma-glutamyl alpha-amino acid
- Glutamine or derivatives
- N-acyl-l-alpha-amino acid
- N-acyl-alpha-amino acid
- N-acyl-alpha amino acid or derivatives
- Alanine or derivatives
- Alpha-amino acid
- Alpha-amino acid or derivatives
- L-alpha-amino acid
- Dicarboxylic acid or derivatives
- Fatty amide
- N-acyl-amine
- Fatty acyl
- Fatty acid
- Amino acid or derivatives
- Carboxamide group
- Amino acid
- Secondary carboxylic acid amide
- Carboxylic acid
- Hydrocarbon derivative
- Primary aliphatic amine
- Organic oxygen compound
- Organic oxide
- Organopnictogen compound
- Organic nitrogen compound
- Carbonyl group
- Amine
- Primary amine
- Organonitrogen compound
- Organooxygen compound
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00al-9710000000-e2845b73a5e7cd81881f | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-00dj-9332000000-f7980440d2535885b7ab | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00xr-2940000000-948d6e9385d802328937 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-05br-6900000000-5f1480e854581ee71678 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4r-9200000000-58a633696455e707016d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014i-0690000000-92c8a111979eb46fa59f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-06dj-2930000000-1156c919eda2b8e49fae | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0076-9200000000-737e76bca1d968721a2a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000x-9330000000-8674f1f30751437e8bf9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-001i-9300000000-a9c7516230cc6b7166c1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-053u-9100000000-bb369086fe31b624af6c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-056r-1900000000-491b535e4d4e3214438c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000i-9100000000-51f9fd3eeccf33a98c1c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-000f-9000000000-d6a5f92f606e802c70be | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0006248 |
|---|
| FooDB ID | FDB023860 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00055691 |
|---|
| BiGG ID | 1446127 |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 389104 |
|---|
| ChEBI ID | 50619 |
|---|
| PubChem Compound ID | 440103 |
|---|
| Kegg Compound ID | C03740 |
|---|
| YMDB ID | YMDB00691 |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. https://www.ncbi.nlm.nih.gov/pubmed/?term=14213360 | | 2. https://www.ncbi.nlm.nih.gov/pubmed/?term=14494011 | | 3. https://www.ncbi.nlm.nih.gov/pubmed/?term=5867340 | | 4. York MJ, Crossley MJ, Hyslop SJ, Fisher ML, Kuchel PW: gamma-Glutamylcyclotransferase: inhibition by D-beta-aminoglutaryl-L-alanine and analysis of the solvent kinetic isotope effect. Eur J Biochem. 1989 Sep 1;184(1):97-101. | | 5. Takahashi T, Kondo T, Ohno H, Minato S, Ohshima T, Mikuni S, Soda K, Taniguchi N: A spectrophotometric method for the determination of gamma-glutamyl cyclotransferase with alanine dehydrogenase in the presence of anthglutin. Biochem Med Metab Biol. 1987 Dec;38(3):311-6. | | 6. Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. |
|
|---|