| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:58:13 UTC |
|---|
| Update Date | 2016-11-09 01:21:21 UTC |
|---|
| Accession Number | CHEM035664 |
|---|
| Identification |
|---|
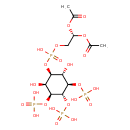
| Common Name | Phosphatidylinositol-3,4,5-trisphosphate |
|---|
| Class | Small Molecule |
|---|
| Description | Phosphatidylinositol (3,4,5)-trisphosphate (PtdIns(3,4,5)P3) commonly abbreviated to PIP3 is the product of the class I phosphoinositide 3-kinases (PI 3-kinases) activity on phosphatidylinositol (4,5)-bisphosphate. PtdIns(3,4,5)P3 is dephophosphorylated by the phosphatase, PTEN on the 3 position and by SHIPs (SH2-containing inositol phosphatase) on the 5' position of the inositol ring.
The PH domain in a number of proteins binds to PtdIns(3,4,5)P3. Such proteins include Akt/PKB, PDK1, Btk1 and ARNO. The generation of PtdIns(3,4,5)P3 at the plasma membrane upon the activation of class I PI 3-kinases causes these proteins to translocate to the plasma membrane and accordingly affects their activity. The PH domain allows binding between PtdIns(3,4,5)P3 and G-protein coupled receptor kinases (GRKs). This enhances the binding of the GRK to the plasma membrane. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| Phosphatidylinositol-3,4,5-trisphosphoric acid | Generator | | 1,2-Diacyl-sn-glycero-3-phospho-(1'-myo-inositol-3',4',5'-bisphosphate) | HMDB | | 1-Phosphatidyl-1D-myo-inositol 3,4,5-trisphosphate | HMDB |
|
|---|
| Chemical Formula | C12H24O22P4 |
|---|
| Average Molecular Mass | 644.201 g/mol |
|---|
| Monoisotopic Mass | 643.971 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | {[(1S,2S,3S,4S,5R,6S)-3-({[2,2-bis(acetyloxy)ethoxy](hydroxy)phosphoryl}oxy)-2,4-dihydroxy-5,6-bis(phosphonooxy)cyclohexyl]oxy}phosphonic acid |
|---|
| Traditional Name | [(1S,2S,3S,4S,5R,6S)-3-{[2,2-bis(acetyloxy)ethoxy(hydroxy)phosphoryl]oxy}-2,4-dihydroxy-5,6-bis(phosphonooxy)cyclohexyl]oxyphosphonic acid |
|---|
| SMILES | CC(=O)OC(COP(O)(=O)O[C@@H]1[C@H](O)[C@H](OP(O)(O)=O)[C@@H](OP(O)(O)=O)[C@H](OP(O)(O)=O)[C@H]1O)OC(C)=O |
|---|
| InChI Identifier | InChI=1S/C12H24O22P4/c1-4(13)29-6(30-5(2)14)3-28-38(26,27)34-9-7(15)10(31-35(17,18)19)12(33-37(23,24)25)11(8(9)16)32-36(20,21)22/h6-12,15-16H,3H2,1-2H3,(H,26,27)(H2,17,18,19)(H2,20,21,22)(H2,23,24,25)/t7-,8-,9-,10+,11-,12-/m0/s1 |
|---|
| InChI Key | RQQIRMLGKSPXSE-UQPICLANSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as inositol phosphates. Inositol phosphates are compounds containing a phosphate group attached to an inositol (or cyclohexanehexol) moiety. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Alcohols and polyols |
|---|
| Direct Parent | Inositol phosphates |
|---|
| Alternative Parents | |
|---|
| Substituents | - Inositol phosphate
- Cyclohexanol
- Acylal
- Dialkyl phosphate
- Monoalkyl phosphate
- Dicarboxylic acid or derivatives
- Organic phosphoric acid derivative
- Phosphoric acid ester
- Alkyl phosphate
- Carboxylic acid ester
- Secondary alcohol
- Acetal
- Carboxylic acid derivative
- Hydrocarbon derivative
- Organic oxide
- Carbonyl group
- Aliphatic homomonocyclic compound
|
|---|
| Molecular Framework | Aliphatic homomonocyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0002-9221350000-b16be7ebadf266750598 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_7) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_8) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_9) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_10) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_11) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_12) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_13) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_14) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_15) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_16) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_17) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_18) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0ika-9200563000-9a60ed0c3fcbf50167f3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03e9-9110533000-fc66e6a74addc319984e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0w29-9204410000-ff2fba934c963beb20a6 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-052f-9000213000-32b5b2c505d6a6a73bd7 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9001211000-a6daae7f68ecbcc9ba34 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9000000000-f50c048414e579ad0a7e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0006-0000794000-d2b55615db3cd4aeead3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-005a-7000911000-7e41bb115a7c12f0d6e3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004l-9100160000-a155ea0d71acc5294fb4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0000092000-8e9c207b385bcdcbaa0c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-1000291000-137cfeaa77a7642499fa | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0006-9000000000-83fc05b0f4b5c6b39135 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0004249 |
|---|
| FooDB ID | FDB023351 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | 47125 |
|---|
| BioCyc ID | PHOSPHATIDYL-INOSITOL-345-TRISPHOSPHAT |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 34980938 |
|---|
| ChEBI ID | 16618 |
|---|
| PubChem Compound ID | Not Available |
|---|
| Kegg Compound ID | C05981 |
|---|
| YMDB ID | YMDB16180 |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Gurung R, Tan A, Ooms LM, McGrath MJ, Huysmans RD, Munday AD, Prescott M, Whisstock JC, Mitchell CA: Identification of a novel domain in two mammalian inositol-polyphosphate 5-phosphatases that mediates membrane ruffle localization. The inositol 5-phosphatase skip localizes to the endoplasmic reticulum and translocates to membrane ruffles following epidermal growth factor stimulation. J Biol Chem. 2003 Mar 28;278(13):11376-85. Epub 2003 Jan 20. | | 2. Ijuin T, Takenawa T: SKIP negatively regulates insulin-induced GLUT4 translocation and membrane ruffle formation. Mol Cell Biol. 2003 Feb;23(4):1209-20. | | 3. Radu A, Neubauer V, Akagi T, Hanafusa H, Georgescu MM: PTEN induces cell cycle arrest by decreasing the level and nuclear localization of cyclin D1. Mol Cell Biol. 2003 Sep;23(17):6139-49. | | 4. Maxwell MJ, Yuan Y, Anderson KE, Hibbs ML, Salem HH, Jackson SP: SHIP1 and Lyn Kinase Negatively Regulate Integrin alpha IIb beta 3 signaling in platelets. J Biol Chem. 2004 Jul 30;279(31):32196-204. Epub 2004 May 27. | | 5. Liu Q, Dumont DJ: Molecular cloning and chromosomal localization in human and mouse of the SH2-containing inositol phosphatase, INPP5D (SHIP). Amgen EST Program. Genomics. 1997 Jan 1;39(1):109-12. |
|
|---|