| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:33:33 UTC |
|---|
| Update Date | 2016-11-09 01:21:16 UTC |
|---|
| Accession Number | CHEM035187 |
|---|
| Identification |
|---|
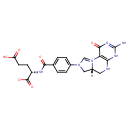
| Common Name | 5,10-Methenyltetrahydrofolic acid |
|---|
| Class | Small Molecule |
|---|
| Description | 5,10-5,10-5,10-methenyltetrahydrofolic acid, also known as 5,10-5,10-methenyltetrahydrofolic acid or 5,10-methenyl-THF, belongs to the class of organic compounds known as tetrahydrofolic acids. These are heterocyclic compounds based on the 5,6,7,8-tetrahydropteroic acid skeleton conjugated with at least one L-glutamic acid unit. 5,10-5,10-5,10-methenyltetrahydrofolic acid is possibly soluble (in water) and a moderately basic compound (based on its pKa). 5,10-5,10-5,10-methenyltetrahydrofolic acid exists in all living species, ranging from bacteria to humans. 5,10-5,10-5,10-methenyltetrahydrofolic acid participates in a number of enzymatic reactions, within cattle. In particular, 5,10-5,10-5,10-methenyltetrahydrofolic acid can be biosynthesized from 10-formyltetrahydrofolate; which is catalyzed by the enzyme C-1-tetrahydrofolate synthase, cytoplasmic. In addition, 5,10-5,10-5,10-methenyltetrahydrofolic acid can be biosynthesized from N5-formyl-THF; which is mediated by the enzyme 5-formyltetrahydrofolate cyclo-ligase. In cattle, 5,10-5,10-methenyltetrahydrofolic acid is involved in the metabolic pathway called the folate metabolism pathway. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 5,10-Methenyltetrahydrofolate | Generator | | 5,10-Methenyl-THF | HMDB | | Anhydro-leucovorin | HMDB | | Anhydro-leucovorin a | HMDB | | Anhydroleucovorin | HMDB | | Anhydroleucovorin a | HMDB | | CH-THF | HMDB | | Methenyl-tetrahydrofolate | HMDB | | Methenyl-THF | HMDB | | Methenyltetrahydrofolate | HMDB | | Methenyltetrahydrofolic acid | HMDB | | N5-N10-CH-THF | HMDB | | N5-N10-Methenyltetrahydrofolate | HMDB | | 5,10-Methenyltetrahydropteroylglutamate | HMDB | | N5,N10-Methenyl tetrahydrofolate | HMDB |
|
|---|
| Chemical Formula | C20H21N7O6 |
|---|
| Average Molecular Mass | 455.424 g/mol |
|---|
| Monoisotopic Mass | 455.155 g/mol |
|---|
| CAS Registry Number | 7444-29-3 |
|---|
| IUPAC Name | (6aR)-8-(4-{[(1S)-1,3-dicarboxypropyl]-C-hydroxycarbonimidoyl}phenyl)-1-hydroxy-3-imino-3H,4H,5H,6H,6aH,7H-8lambda5-imidazo[1,5-f]pteridin-8-ylium |
|---|
| Traditional Name | (6aR)-8-(4-{[(1S)-1,3-dicarboxypropyl]-C-hydroxycarbonimidoyl}phenyl)-1-hydroxy-3-imino-4H,5H,6H,6aH,7H-8lambda5-imidazo[1,5-f]pteridin-8-ylium |
|---|
| SMILES | [H][C@@]12CN(C=[N+]1C1=C(NC2)NC(N)=NC1=O)C1=CC=C(C=C1)C(=O)N[C@@H](CCC(O)=O)C([O-])=O |
|---|
| InChI Identifier | InChI=1S/C20H21N7O6/c21-20-24-16-15(18(31)25-20)27-9-26(8-12(27)7-22-16)11-3-1-10(2-4-11)17(30)23-13(19(32)33)5-6-14(28)29/h1-4,9,12-13H,5-8H2,(H6-,21,22,23,24,25,28,29,30,31,32,33)/t12-,13+/m1/s1 |
|---|
| InChI Key | MEANFMOQMXYMCT-OLZOCXBDSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as tetrahydrofolic acids. These are heterocyclic compounds based on the 5,6,7,8-tetrahydropteroic acid skeleton conjugated with at least one L-glutamic acid unit. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Pteridines and derivatives |
|---|
| Sub Class | Pterins and derivatives |
|---|
| Direct Parent | Tetrahydrofolic acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Tetrahydrofolic acid
- Glutamic acid or derivatives
- N-acyl-alpha amino acid or derivatives
- N-acyl-alpha-amino acid
- Hippuric acid
- Hippuric acid or derivatives
- Alpha-amino acid or derivatives
- Benzamide
- Benzoic acid or derivatives
- Aniline or substituted anilines
- Benzoyl
- Aminopyrimidine
- Pyrimidone
- Secondary aliphatic/aromatic amine
- Monocyclic benzene moiety
- Pyrimidine
- Dicarboxylic acid or derivatives
- Benzenoid
- Vinylogous amide
- Heteroaromatic compound
- 2-imidazoline
- Carboxamide group
- Amino acid
- Secondary carboxylic acid amide
- Carboxylic acid salt
- Amino acid or derivatives
- Amidine
- Azacycle
- Carboxylic acid amidine
- Organic 1,3-dipolar compound
- Carboxylic acid derivative
- Propargyl-type 1,3-dipolar organic compound
- Carboxylic acid
- Secondary amine
- Organic oxygen compound
- Amine
- Carbonyl group
- Organic salt
- Organic nitrogen compound
- Organopnictogen compound
- Organic oxide
- Organic zwitterion
- Hydrocarbon derivative
- Primary amine
- Organooxygen compound
- Organonitrogen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-03di-2408900000-90c26f197ed01948843b | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-0230-9506630000-d9b025a3f36832b17d97 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4i-0000900000-9a74d30be444557240b0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4i-0000900000-87ffec278a0d552070d3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-056r-9210100000-8e44a81962b4227dbc32 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0000900000-6e21d81553cf048dcee3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0udi-2000900000-e3e2170131566f2c34e9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9300100000-8e0fa01b9b99e8b1fea8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4i-0003900000-a08e1d1cb1452492cc40 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0bt9-2109100000-a099f262b4e99916ab71 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-053r-0396000000-4a34a1ca0c2780791f62 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0f79-0004900000-b7d25e9a9f2b33373222 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001u-4129500000-674f21af43f4a24fc9aa | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0f9y-9643100000-a3b87dfc44080a520c03 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0001354 |
|---|
| FooDB ID | FDB022573 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | 34999 |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | 6185 |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | 5,10-Methenyltetrahydrofolate |
|---|
| Chemspider ID | 559356 |
|---|
| ChEBI ID | 15636 |
|---|
| PubChem Compound ID | 135450599 |
|---|
| Kegg Compound ID | C00445 |
|---|
| YMDB ID | YMDB00176 |
|---|
| ECMDB ID | ECMDB24067 |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Field MS, Anderson DD, Stover PJ: Mthfs is an Essential Gene in Mice and a Component of the Purinosome. Front Genet. 2011 Jun 20;2:36. doi: 10.3389/fgene.2011.00036. eCollection 2011. | | 2. Oberpichler I, Pierik AJ, Wesslowski J, Pokorny R, Rosen R, Vugman M, Zhang F, Neubauer O, Ron EZ, Batschauer A, Lamparter T: A photolyase-like protein from Agrobacterium tumefaciens with an iron-sulfur cluster. PLoS One. 2011;6(10):e26775. doi: 10.1371/journal.pone.0026775. Epub 2011 Oct 31. | | 3. Telegina TA, Liudnikova TA, Zemskova IuL, Sviridov EA, Kritskii MS: [Tolerance of 5,10-methenyltetrahydrofolate to ultraviolet radiation]. Prikl Biokhim Mikrobiol. 2005 May-Jun;41(3):315-23. | | 4. Eadsforth TC, Cameron S, Hunter WN: The crystal structure of Leishmania major N(5),N(10)-methylenetetrahydrofolate dehydrogenase/cyclohydrolase and assessment of a potential drug target. Mol Biochem Parasitol. 2012 Feb;181(2):178-85. doi: 10.1016/j.molbiopara.2011.11.004. Epub 2011 Nov 15. | | 5. Lin CJ, Wen MJ, Hung YJ, Pei D, Kuo SW, Hsieh CH: The impact of 5,10-methenyltetrahydrofolate synthetase polymorphism on diabetic nephropathy in the Taiwanese population. Genet Test Mol Biomarkers. 2012 Feb;16(2):142-5. doi: 10.1089/gtmb.2011.0050. Epub 2011 Sep 6. | | 6. Field MS, Szebenyi DM, Stover PJ: Regulation of de novo purine biosynthesis by methenyltetrahydrofolate synthetase in neuroblastoma. J Biol Chem. 2006 Feb 17;281(7):4215-21. Epub 2005 Dec 19. | | 7. Kruszyna L, Lianeri M, Rydzanicz M, Gajecka M, Szyfter K, Jagodzinski PP: Polymorphic variants of folate metabolism genes and the risk of laryngeal cancer. Mol Biol Rep. 2010 Jan;37(1):241-7. doi: 10.1007/s11033-009-9643-y. Epub 2009 Aug 1. | | 8. Moldt J, Pokorny R, Orth C, Linne U, Geisselbrecht Y, Marahiel MA, Essen LO, Batschauer A: Photoreduction of the folate cofactor in members of the photolyase family. J Biol Chem. 2009 Aug 7;284(32):21670-83. doi: 10.1074/jbc.M109.018697. Epub 2009 Jun 16. | | 9. Ogwang S, Nguyen HT, Sherman M, Bajaksouzian S, Jacobs MR, Boom WH, Zhang GF, Nguyen L: Bacterial conversion of folinic acid is required for antifolate resistance. J Biol Chem. 2011 Apr 29;286(17):15377-90. doi: 10.1074/jbc.M111.231076. Epub 2011 Mar 3. | | 10. Watkins D, Rosenblatt DS: Update and new concepts in vitamin responsive disorders of folate transport and metabolism. J Inherit Metab Dis. 2012 Jul;35(4):665-70. doi: 10.1007/s10545-011-9418-1. Epub 2011 Nov 23. | | 11. Kariluoto S, Edelmann M, Herranen M, Lampi AM, Shmelev A, Salovaara H, Korhola M, Piironen V: Production of folate by bacteria isolated from oat bran. Int J Food Microbiol. 2010 Sep 30;143(1-2):41-7. doi: 10.1016/j.ijfoodmicro.2010.07.026. Epub 2010 Aug 11. | | 12. Selby CP, Sancar A: The second chromophore in Drosophila photolyase/cryptochrome family photoreceptors. Biochemistry. 2012 Jan 10;51(1):167-71. doi: 10.1021/bi201536w. Epub 2011 Dec 27. | | 13. Jahansouz H, Scherubel DM, Himes RH: Formylation of tetrahydrofolate by formyl phosphate. FEBS Lett. 1990 Mar 26;262(2):366-8. | | 14. Tolley M, Bickford L, Clare K, Johann TW: Investigations of amino acids in the ATP binding site of 5,10-methenyltetrahydrofolate synthetase. Protein J. 2012 Aug;31(6):519-28. doi: 10.1007/s10930-012-9428-3. | | 15. Wijaya IM, Zhang Y, Iwata T, Yamamoto J, Hitomi K, Iwai S, Getzoff ED, Kandori H: Detection of distinct alpha-helical rearrangements of cyclobutane pyrimidine dimer photolyase upon substrate binding by Fourier transform infrared spectroscopy. Biochemistry. 2013 Feb 12;52(6):1019-27. doi: 10.1021/bi3016179. Epub 2013 Jan 30. | | 16. Upadhyay V, Demmer U, Warkentin E, Moll J, Shima S, Ermler U: Structure and catalytic mechanism of N(5),N(10)-methenyl-tetrahydromethanopterin cyclohydrolase. Biochemistry. 2012 Oct 23;51(42):8435-43. doi: 10.1021/bi300777k. Epub 2012 Oct 8. | | 17. Kirsch SH, Knapp JP, Herrmann W, Obeid R: Quantification of key folate forms in serum using stable-isotope dilution ultra performance liquid chromatography-tandem mass spectrometry. J Chromatogr B Analyt Technol Biomed Life Sci. 2010 Jan 1;878(1):68-75. doi: 10.1016/j.jchromb.2009.11.021. | | 18. Knock E, Deng L, Krupenko N, Mohan RD, Wu Q, Leclerc D, Gupta S, Elmore CL, Kruger W, Tini M, Rozen R: Susceptibility to intestinal tumorigenesis in folate-deficient mice may be influenced by variation in one-carbon metabolism and DNA repair. J Nutr Biochem. 2011 Nov;22(11):1022-9. doi: 10.1016/j.jnutbio.2010.07.015. Epub 2010 Dec 28. |
|
|---|