| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:31:38 UTC |
|---|
| Update Date | 2016-11-09 01:21:15 UTC |
|---|
| Accession Number | CHEM035147 |
|---|
| Identification |
|---|
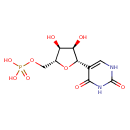
| Common Name | Pseudouridine 5'-phosphate |
|---|
| Class | Small Molecule |
|---|
| Description | A C-nucleoside phosphate consisting of pseudouridine having a monophosphate group at the 5'-position. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| (1S)-1,4-Anhydro-1-(2,4-dioxo-1,2,3,4-tetrahydropyrimidin-5-yl)-5-O-phosphono-D-ribitol | ChEBI | | 5-(5-O-Phosphono-beta-D-ribofuranosyl)pyrimidine-2,4(1H,3H)-dione | ChEBI | | 5-(5-O-Phosphono-b-D-ribofuranosyl)pyrimidine-2,4(1H,3H)-dione | Generator | | 5-(5-O-Phosphono-β-D-ribofuranosyl)pyrimidine-2,4(1H,3H)-dione | Generator | | Pseudouridine 5'-phosphoric acid | Generator | | Pseudouridylic acid | MeSH, HMDB | | 5-(5-O-Phosphono-beta-D-ribofuranosyl)-2,4(1H,3H)-pyrimidinedione | HMDB | | 5-(5-O-Phosphono-β-D-ribofuranosyl)-2,4(1H,3H)-pyrimidinedione | HMDB | | Pseudouridine 5'-phosphate | HMDB | | Pseudouridine 5’-phosphate | HMDB | | Pseudouridine monophosphate | HMDB |
|
|---|
| Chemical Formula | C9H13N2O9P |
|---|
| Average Molecular Mass | 324.181 g/mol |
|---|
| Monoisotopic Mass | 324.036 g/mol |
|---|
| CAS Registry Number | 1157-60-4 |
|---|
| IUPAC Name | {[(2R,3S,4R,5S)-5-(2,4-dioxo-1,2,3,4-tetrahydropyrimidin-5-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}phosphonic acid |
|---|
| Traditional Name | pseudouridine 5'-phosphate |
|---|
| SMILES | O[C@H]1[C@@H](O)[C@@H](O[C@@H]1COP(O)(O)=O)C1=CNC(=O)NC1=O |
|---|
| InChI Identifier | InChI=1S/C9H13N2O9P/c12-5-4(2-19-21(16,17)18)20-7(6(5)13)3-1-10-9(15)11-8(3)14/h1,4-7,12-13H,2H2,(H2,16,17,18)(H2,10,11,14,15)/t4-,5-,6-,7+/m1/s1 |
|---|
| InChI Key | MOBMOJGXNHLLIR-GBNDHIKLSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as pentose phosphates. These are carbohydrate derivatives containing a pentose substituted by one or more phosphate groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Pentose phosphates |
|---|
| Alternative Parents | |
|---|
| Substituents | - Pentose phosphate
- Pentose-5-phosphate
- C-glycosyl compound
- Glycosyl compound
- Monosaccharide phosphate
- Pyrimidone
- Monoalkyl phosphate
- Hydropyrimidine
- Organic phosphoric acid derivative
- Phosphoric acid ester
- Pyrimidine
- Alkyl phosphate
- Tetrahydrofuran
- Vinylogous amide
- Heteroaromatic compound
- Secondary alcohol
- Urea
- 1,2-diol
- Lactam
- Dialkyl ether
- Azacycle
- Ether
- Organoheterocyclic compound
- Oxacycle
- Organonitrogen compound
- Alcohol
- Organic oxide
- Organopnictogen compound
- Hydrocarbon derivative
- Organic nitrogen compound
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0092-7921000000-3b484f48c79da75d22b9 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-0zor-5984100000-c1d4654dc77f1a2defc3 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-004i-0669000000-0b19df9acf8df54d0c28 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-004j-4493000000-38400e5991956cb67184 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000l-3900000000-ebfc1c181ae3a767f1d5 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0096-9022000000-61eca04124385371e070 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004l-9000000000-d8f1a9a373cb543c5a50 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9000000000-7662d761a5f16134b340 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-00fr-9007000000-706b8bb8bcb6e1f088ab | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9000000000-a5e502a2627af2048a1f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9000000000-a5e502a2627af2048a1f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-004i-0191000000-d8971429f6e3bcedb1b2 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-056r-0960000000-db21c37dfcf9636eeeb4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03di-2900000000-8e2c2edc3141326a4fb1 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB03829 |
|---|
| HMDB ID | HMDB0001271 |
|---|
| FooDB ID | FDB022525 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | PSEUDOURIDINE-5-P |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 388535 |
|---|
| ChEBI ID | 18116 |
|---|
| PubChem Compound ID | 439424 |
|---|
| Kegg Compound ID | C01168 |
|---|
| YMDB ID | YMDB00780 |
|---|
| ECMDB ID | ECMDB01271 |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Charette M, Gray MW: Pseudouridine in RNA: what, where, how, and why. IUBMB Life. 2000 May;49(5):341-51. | | 2. Brunk E, Sahoo S, Zielinski DC, Altunkaya A, Drager A, Mih N, Gatto F, Nilsson A, Preciat Gonzalez GA, Aurich MK, Prlic A, Sastry A, Danielsdottir AD, Heinken A, Noronha A, Rose PW, Burley SK, Fleming RMT, Nielsen J, Thiele I, Palsson BO: Recon3D enables a three-dimensional view of gene variation in human metabolism. Nat Biotechnol. 2018 Mar;36(3):272-281. doi: 10.1038/nbt.4072. Epub 2018 Feb 19. |
|
|---|