| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:28:53 UTC |
|---|
| Update Date | 2016-11-09 01:21:15 UTC |
|---|
| Accession Number | CHEM035090 |
|---|
| Identification |
|---|
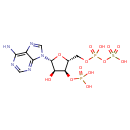
| Common Name | Phosphoadenosine phosphosulfate |
|---|
| Class | Small Molecule |
|---|
| Description | Phosphoadenosine phosphosulfate is a key intermediate in the formation by living cells of sulfate esters of phenols, alcohols, steroids, sulfated polysaccharides, and simple esters, such as choline sulfate. It is formed from sulfate ion and ATP in a two-step process. This compound also is an important step in the process of sulfur fixation in plants and microorganisms. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 3'-Phosphoadenosine 5'-phosphosulfate | ChEBI | | 3'-Phosphoadenylyl sulfate | ChEBI | | PAPS | ChEBI | | 3'-Phospho-5'-adenylyl sulfate | Kegg | | 3'-Phosphoadenosine 5'-phosphosulfuric acid | Generator | | 3'-Phosphoadenosine 5'-phosphosulphate | Generator | | 3'-Phosphoadenosine 5'-phosphosulphuric acid | Generator | | 3'-Phosphoadenylyl sulfuric acid | Generator | | 3'-Phosphoadenylyl sulphate | Generator | | 3'-Phosphoadenylyl sulphuric acid | Generator | | 3'-Phospho-5'-adenylyl sulfuric acid | Generator | | 3'-Phospho-5'-adenylyl sulphate | Generator | | 3'-Phospho-5'-adenylyl sulphuric acid | Generator | | Phosphoadenosine phosphosulfuric acid | Generator | | Phosphoadenosine phosphosulphate | Generator | | Phosphoadenosine phosphosulphuric acid | Generator | | 3'-Phosphoadenosine-5'-phosphosulfate | HMDB | | 3'-Phosphoadenosine-5'-phosphosulphate | HMDB | | 3'-Phosphoadenylyl-sulfate | HMDB | | 3'-Phosphoadenylyl-sulphate | HMDB | | 5-Phosphoadenosine 3-phosphosulfate | HMDB | | 5-Phosphoadenosine 3-phosphosulphate | HMDB | | Adenosine 3' phosphate 5' phosphosulfate | HMDB | | Adenosine-3'-phosphate-5'-phosphosulfate | HMDB | | Phosphosulfate, phosphoadenosine | HMDB | | 3’-Phosphoadenosine 5’-phosphosulfate | HMDB | | 3’-Phosphoadenosine 5’-phosphosulphate | HMDB | | 3’-Phosphoadenylyl sulfate | HMDB | | 3’-Phosphoadenylyl sulphate | HMDB | | 5'-Adenylyl sulfate 3'-phosphate | HMDB | | 5'-Adenylyl sulphate 3'-phosphate | HMDB | | 5’-Adenylyl sulfate 3’-phosphate | HMDB | | 5’-Adenylyl sulphate 3’-phosphate | HMDB | | Adenosine 3'-phosphate 5'-phosphosulfate | HMDB | | Adenosine 3'-phosphate 5'-phosphosulphate | HMDB | | Adenosine 3'-phosphate 5'-sulfatophosphate | HMDB | | Adenosine 3’-phosphate 5’-phosphosulfate | HMDB | | Adenosine 3’-phosphate 5’-phosphosulphate | HMDB | | Adenosine 3’-phosphate 5’-sulfatophosphate | HMDB | | Adenosine 5'-phosphosulfate 3'-phosphate | HMDB | | Adenosine 5'-phosphosulphate 3'-phosphate | HMDB | | Adenosine 5’-phosphosulfate 3’-phosphate | HMDB | | Adenosine 5’-phosphosulphate 3’-phosphate | HMDB |
|
|---|
| Chemical Formula | C10H15N5O13P2S |
|---|
| Average Molecular Mass | 507.264 g/mol |
|---|
| Monoisotopic Mass | 506.986 g/mol |
|---|
| CAS Registry Number | 482-67-7 |
|---|
| IUPAC Name | [({[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-4-hydroxy-3-(phosphonooxy)oxolan-2-yl]methoxy}(hydroxy)phosphoryl)oxy]sulfonic acid |
|---|
| Traditional Name | 3'-phosphoadenylyl sulfate |
|---|
| SMILES | NC1=NC=NC2=C1N=CN2[C@@H]1O[C@H](COP(O)(=O)OS(O)(=O)=O)[C@@H](OP(O)(O)=O)[C@H]1O |
|---|
| InChI Identifier | InChI=1S/C10H15N5O13P2S/c11-8-5-9(13-2-12-8)15(3-14-5)10-6(16)7(27-29(17,18)19)4(26-10)1-25-30(20,21)28-31(22,23)24/h2-4,6-7,10,16H,1H2,(H,20,21)(H2,11,12,13)(H2,17,18,19)(H,22,23,24)/t4-,6-,7-,10-/m1/s1 |
|---|
| InChI Key | GACDQMDRPRGCTN-KQYNXXCUSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as purine ribonucleoside 3',5'-bisphosphates. These are purine ribobucleotides with one phosphate group attached to 3' and 5' hydroxyl groups of the ribose moiety. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Nucleosides, nucleotides, and analogues |
|---|
| Class | Purine nucleotides |
|---|
| Sub Class | Purine ribonucleotides |
|---|
| Direct Parent | Purine ribonucleoside 3',5'-bisphosphates |
|---|
| Alternative Parents | |
|---|
| Substituents | - Purine ribonucleoside 3',5'-bisphosphate
- Purine ribonucleoside monophosphate
- Ribonucleoside 3'-phosphate
- Pentose phosphate
- Pentose-5-phosphate
- Glycosyl compound
- N-glycosyl compound
- 6-aminopurine
- Monosaccharide phosphate
- Pentose monosaccharide
- Imidazopyrimidine
- Purine
- Aminopyrimidine
- Monoalkyl phosphate
- Monosaccharide
- N-substituted imidazole
- Organic phosphoric acid derivative
- Phosphoric acid ester
- Imidolactam
- Alkyl phosphate
- Pyrimidine
- Tetrahydrofuran
- Organic sulfuric acid or derivatives
- Azole
- Imidazole
- Heteroaromatic compound
- Secondary alcohol
- Organoheterocyclic compound
- Azacycle
- Oxacycle
- Organooxygen compound
- Organic nitrogen compound
- Hydrocarbon derivative
- Alcohol
- Organic oxide
- Organic oxygen compound
- Primary amine
- Amine
- Organonitrogen compound
- Organopnictogen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0002-8930200000-a2821e129c20bee87c33 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-0002-9441110000-619798b0c3242743c13a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-2901410000-afa8a71104e69c842868 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0900200000-8f9317ccb6340a4317c4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-1900000000-e2c92e775605a11d728e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a7i-4900240000-500302d146b6d356a0b8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0059-5900100000-07e0047589df3afabbbd | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9500000000-517ff9b08c8f8999b922 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4i-0000090000-ba006402f6f0af41d779 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-01ri-0402920000-7d549a97dd2276b8b754 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-01qi-0439100000-94b8e2004e4fb012c91d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-0900000000-63709bad7395f9f33acc | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-0900000000-94a8fdc8360cf2577ca6 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-003s-8900000000-0b1152e8f0b7373e9349 | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB02902 |
|---|
| HMDB ID | HMDB0001134 |
|---|
| FooDB ID | FDB022445 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00007446 |
|---|
| BiGG ID | 33679 |
|---|
| BioCyc ID | PAPS |
|---|
| METLIN ID | 6028 |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | 3'-Phosphoadenosine-5'-phosphosulfate |
|---|
| Chemspider ID | 9799 |
|---|
| ChEBI ID | 17980 |
|---|
| PubChem Compound ID | 10214 |
|---|
| Kegg Compound ID | C00053 |
|---|
| YMDB ID | YMDB00120 |
|---|
| ECMDB ID | ECMDB01134 |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Lin, Chun-Hung; Shen, Gwo-Jenn; Garcia-Junceda, Eduardo; Wong, Chi-Huey. Enzymic Synthesis and Regeneration of 3'-Phosphoadenosine 5'-Phosphosulfate (PAPS) for Regioselective Sulfation of Oligosaccharides. J. Am. Chem. Soc., 1995, 117 (30), pp 8031-8032 | | 2. Emmi L, Bergamini C, Spinelli A, Liotta F, Marchione T, Caldini A, Fanelli A, De Cristofaro MT, Dal Pozzo G: Possible pathogenetic role of activated platelets in the primary antiphospholipid syndrome involving the central nervous system. Ann N Y Acad Sci. 1997 Aug 14;823:188-200. | | 3. Fanelli A, Bergamini C, Rapi S, Caldini A, Spinelli A, Buggiani A, Emmi L: Flow cytometric detection of circulating activated platelets in primary antiphospholipid syndrome. Correlation with thrombocytopenia and anticardiolipin antibodies. Lupus. 1997;6(3):261-7. | | 4. Joseph JE, Harrison P, Mackie IJ, Isenberg DA, Machin SJ: Increased circulating platelet-leucocyte complexes and platelet activation in patients with antiphospholipid syndrome, systemic lupus erythematosus and rheumatoid arthritis. Br J Haematol. 2001 Nov;115(2):451-9. | | 5. Suarez IM, Diaz RA, Aguayo Canela D, Pujol de la Llave E: Correction of severe thrombocytopenia with chloroquine in the primary antiphospholipid syndrome. Lupus. 1996 Feb;5(1):81-3. | | 6. Khoo BY, Sit KH, Wong KP: Does PAPS generation determine the overall sulfate conjugation in human platelets? Life Sci. 1988;42(23):2389-95. | | 7. Wong KP, Khoo BY, Sit KH: Biosynthesis of PAPS in vitro by human liver. Measurement by two independent assay procedures. Biochem Pharmacol. 1991 Jan 1;41(1):63-9. | | 8. Cappiello M, Franchi M, Rane A, Pacifici GM: Sulphotransferase and its substrate: adenosine-3'-phosphate-5'-phosphosulphate in human fetal liver and placenta. Dev Pharmacol Ther. 1990;14(1):62-5. | | 9. Cappiello M, Franchi M, Giuliani L, Pacifici GM: Distribution of 2-naphthol sulphotransferase and its endogenous substrate adenosine 3'-phosphate 5'-phosphosulphate in human tissues. Eur J Clin Pharmacol. 1989;37(3):317-20. | | 10. Carlier M, Squifflet JP, Pirson Y, Gribomont B, Alexandre GP: Maximal hydration during anesthesia increases pulmonary arterial pressures and improves early function of human renal transplants. Transplantation. 1982 Oct;34(4):201-4. |
|
|---|