| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:28:22 UTC |
|---|
| Update Date | 2016-11-09 01:21:14 UTC |
|---|
| Accession Number | CHEM035077 |
|---|
| Identification |
|---|
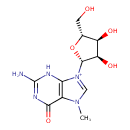
| Common Name | 7-Methylguanosine |
|---|
| Class | Small Molecule |
|---|
| Description | A positively charged methylguanosine in which a single methyl substituent is located at position 7. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| G | ChEBI | | m7g | ChEBI | | N(7)-Methylguanosine | ChEBI | | 2-amino-6,9-dihydro-7-Methyl-6-oxo-9-beta-D-ribofuranosyl-1H-purinium | HMDB | | 2-amino-6,9-dihydro-7-Methyl-6-oxo-9-beta-delta-ribofuranosyl-1H-purinium | HMDB | | N(2)-Methylguanosine | MeSH, HMDB | | 2-Methylguanosine | MeSH, HMDB |
|
|---|
| Chemical Formula | C11H16N5O5 |
|---|
| Average Molecular Mass | 298.275 g/mol |
|---|
| Monoisotopic Mass | 298.115 g/mol |
|---|
| CAS Registry Number | 20244-86-4 |
|---|
| IUPAC Name | 2-amino-9-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]-7-methyl-6-oxo-6,7-dihydro-3H-9lambda5-purin-9-ylium |
|---|
| Traditional Name | 2-amino-9-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]-7-methyl-6-oxo-3H-9lambda5-purin-9-ylium |
|---|
| SMILES | CN1C=[N+]([C@@H]2O[C@H](CO)[C@@H](O)[C@H]2O)C2=C1C(=O)N=C(N)N2 |
|---|
| InChI Identifier | InChI=1S/C11H15N5O5/c1-15-3-16(8-5(15)9(20)14-11(12)13-8)10-7(19)6(18)4(2-17)21-10/h3-4,6-7,10,17-19H,2H2,1H3,(H2-,12,13,14,20)/p+1/t4-,6-,7-,10-/m1/s1 |
|---|
| InChI Key | OGHAROSJZRTIOK-KQYNXXCUSA-O |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as purine nucleosides. Purine nucleosides are compounds comprising a purine base attached to a ribosyl or deoxyribosyl moiety. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Nucleosides, nucleotides, and analogues |
|---|
| Class | Purine nucleosides |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Purine nucleosides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Purine nucleoside
- Glycosyl compound
- N-glycosyl compound
- 6-oxopurine
- Hypoxanthine
- Pentose monosaccharide
- Imidazopyrimidine
- Purine
- Aminopyrimidine
- Pyrimidone
- Monosaccharide
- N-substituted imidazole
- Pyrimidine
- Azole
- Imidazole
- Heteroaromatic compound
- Vinylogous amide
- Tetrahydrofuran
- Secondary alcohol
- Organoheterocyclic compound
- Oxacycle
- Azacycle
- Primary alcohol
- Organic oxygen compound
- Organonitrogen compound
- Organooxygen compound
- Amine
- Alcohol
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Organic nitrogen compound
- Primary amine
- Organic cation
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-004l-7490000000-25f4400703ded15175af | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (3 TMS) - 70eV, Positive | splash10-0a71-9602300000-28a0b9e22a4d9107eecc | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-014i-0920000000-f380f5b5ae1c6d006622 | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-014i-0900000000-4b027f424e6fda6ac5e7 | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-00kb-0900000000-f6e41ae90d0d537e5c37 | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-014i-0900000000-ef3a6eca6ca4a5c643d3 | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-014i-0900000000-1331e6ba04657b4693b4 | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-014i-0920000000-a30f275ae277e41809eb | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-014i-0900000000-2dd6635ba6023cecb388 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0002-0090000000-2168dc01c81274bd0a36 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-3090000000-139798febce24d971377 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-006x-9720000000-3ec4fc5a6820e1393195 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0002-0090000000-8158855cfe3f6155e791 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000t-1490000000-9d89b248d7c16e9713a3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9110000000-0d0610ddbc1f29ed8c2f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-014i-0930000000-a027b5082f87f83617de | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-014i-0900000000-69707776c4ea39129157 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-014i-1900000000-4043fc0f3abc38c203e1 | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 2D NMR | [1H,13C] 2D NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB03493 |
|---|
| HMDB ID | HMDB0001107 |
|---|
| FooDB ID | FDB022428 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00051958 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | CPD0-1041 |
|---|
| METLIN ID | 6008 |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | 7-Methylguanosine |
|---|
| Chemspider ID | 393054 |
|---|
| ChEBI ID | 20794 |
|---|
| PubChem Compound ID | 445404 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. https://www.ncbi.nlm.nih.gov/pubmed/?term=22770225 | | 2. Reynaud C, Bruno C, Boullanger P, Grange J, Barbesti S, Niveleau A: Monitoring of urinary excretion of modified nucleosides in cancer patients using a set of six monoclonal antibodies. Cancer Lett. 1992 Jan 31;61(3):255-62. | | 3. Topp H, Sander G, Heller-Schoch G, Schoch G: Determination of 7-methylguanine, N2,N2-dimethylguanosine, and pseudouridine in ultrafiltrated serum of healthy adults by high-performance liquid chromatography. Anal Biochem. 1987 Feb 15;161(1):49-56. | | 4. Tebib JG, Reynaud C, Cedoz JP, Letroublon MC, Niveleau A: Relationship between urinary excretion of modified nucleosides and rheumatoid arthritis process. Br J Rheumatol. 1997 Sep;36(9):990-5. | | 5. Morris GS, Simmonds HA, Davies PM: Use of biological fluids for the rapid diagnosis of potentially lethal inherited disorders of human purine and pyrimidine metabolism. Biomed Chromatogr. 1986 Jun;1(3):109-18. | | 6. Yang TH, Hu ML: Intracellular levels of S-adenosylhomocysteine but not homocysteine are highly correlated to the expression of nm23-H1 and the level of 5-methyldeoxycytidine in human hepatoma cells with different invasion activities. Nutr Cancer. 2006;55(2):224-31. | | 7. Liu Z, Liu S, Xie Z, Blum W, Perrotti D, Paschka P, Klisovic R, Byrd J, Chan KK, Marcucci G: Characterization of in vitro and in vivo hypomethylating effects of decitabine in acute myeloid leukemia by a rapid, specific and sensitive LC-MS/MS method. Nucleic Acids Res. 2007;35(5):e31. Epub 2007 Jan 30. | | 8. Cho SH, Jung BH, Lee SH, Lee WY, Kong G, Chung BC: Direct determination of nucleosides in the urine of patients with breast cancer using column-switching liquid chromatography-tandem mass spectrometry. Biomed Chromatogr. 2006 Nov;20(11):1229-36. |
|
|---|