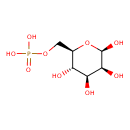| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:27:28 UTC |
|---|
| Update Date | 2016-11-09 01:21:14 UTC |
|---|
| Accession Number | CHEM035063 |
|---|
| Identification |
|---|
| Common Name | Mannose 6-phosphate |
|---|
| Class | Small Molecule |
|---|
| Description | Mannose 6-phosphate, also known as alpha-D-mannose-6-p or mannose 6-phosphate, belongs to the class of organic compounds known as hexose phosphates. These are carbohydrate derivatives containing a hexose substituted by one or more phosphate groups. Mannose 6-phosphate is possibly soluble (in water) and an extremely weak basic (essentially neutral) compound (based on its pKa). Mannose 6-phosphate exists in all eukaryotes, ranging from yeast to humans. Mannose 6-phosphate participates in a number of enzymatic reactions, within cattle. In particular, Mannose 6-phosphate can be converted into fructose 6-phosphate; which is catalyzed by the enzyme mannose 6-phosphate isomerase. In addition, Mannose 6-phosphate can be biosynthesized from D-mannose through the action of the enzyme hexokinase-1. In cattle, mannose 6-phosphate is involved in the metabolic pathway called the fructose and mannose degradation pathway. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| Mannose 6-phosphoric acid | Generator | | alpha-D-Mannose-6-p | HMDB | | alpha-D-Mannose-6-phosphate | HMDB | | alpha-delta-Mannose-6-p | HMDB | | alpha-delta-Mannose-6-phosphate | HMDB | | D-Mannose 6-phosphate | HMDB | | delta-Mannose 6-phosphate | HMDB | | Man-6-p | HMDB | | Mannose-6-phosphate | HMDB | | Mannose-6-phosphate dilithium salt | HMDB | | Mannose-6-phosphate disodium salt | HMDB | | Mannose-6-phosphate sodium salt, (D)-isomer | HMDB |
|
|---|
| Chemical Formula | C6H13O9P |
|---|
| Average Molecular Mass | 260.136 g/mol |
|---|
| Monoisotopic Mass | 260.030 g/mol |
|---|
| CAS Registry Number | 3672-15-9 |
|---|
| IUPAC Name | {[(2R,3S,4S,5S,6R)-3,4,5,6-tetrahydroxyoxan-2-yl]methoxy}phosphonic acid |
|---|
| Traditional Name | mannose 6 phosphate |
|---|
| SMILES | O[C@@H]1O[C@H](COP(O)(O)=O)[C@@H](O)[C@H](O)[C@@H]1O |
|---|
| InChI Identifier | InChI=1S/C6H13O9P/c7-3-2(1-14-16(11,12)13)15-6(10)5(9)4(3)8/h2-10H,1H2,(H2,11,12,13)/t2-,3-,4+,5+,6-/m1/s1 |
|---|
| InChI Key | NBSCHQHZLSJFNQ-RWOPYEJCSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as hexose phosphates. These are carbohydrate derivatives containing a hexose substituted by one or more phosphate groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Hexose phosphates |
|---|
| Alternative Parents | |
|---|
| Substituents | - Hexose phosphate
- Monosaccharide phosphate
- Monoalkyl phosphate
- Organic phosphoric acid derivative
- Alkyl phosphate
- Oxane
- Phosphoric acid ester
- Hemiacetal
- Secondary alcohol
- Oxacycle
- Organoheterocyclic compound
- Polyol
- Hydrocarbon derivative
- Alcohol
- Organic oxide
- Aliphatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aliphatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0005-9740000000-a6e81224fb97751dc2e6 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (4 TMS) - 70eV, Positive | splash10-001i-2533590000-b6a62e882bfffed3184a | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-0007-3490000000-734a88a9b3fd55b3e43f | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-052b-9300000000-2c63b28f70b6b2eb75a1 | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-00l2-9000000000-5406aea843f73d03563c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-0490000000-94ea79ec673f4688b7a0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03dm-5940000000-c649262446626dcb8d6e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0uka-9800000000-aae2cb792906d2dec6cd | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a6r-8290000000-8e20219f94b3fe964e38 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9000000000-b19b64f5cb4124047baf | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9000000000-ecb75a44e3d25affdf31 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a6s-9060000000-b446300429049a22bea4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9000000000-d5aaca13a836cb9f1725 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9000000000-a5e502a2627af2048a1f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03dm-0970000000-40dbaf3d3c72cf65bf47 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-01ot-7900000000-00bc03290b6c0bfb7f84 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000t-9000000000-7a6ff6606f1dbc2f0ae4 | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 2D NMR | [1H,13C] 2D NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0001078 |
|---|
| FooDB ID | FDB022411 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | 34471 |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | 5987 |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Mannose 6-phosphate |
|---|
| Chemspider ID | 388338 |
|---|
| ChEBI ID | 49728 |
|---|
| PubChem Compound ID | 439198 |
|---|
| Kegg Compound ID | C00275 |
|---|
| YMDB ID | YMDB00242 |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Pascual, C.; Herrera, L. Use of permeabilized yeast cells as a system of enzyme immobilization. Its use for the preparation of mannose 6-phosphate. Folia Microbiologica (Prague, Czech Republic) (1981), 26(2), 103-6. | | 2. Beljaars L, Molema G, Weert B, Bonnema H, Olinga P, Groothuis GM, Meijer DK, Poelstra K: Albumin modified with mannose 6-phosphate: A potential carrier for selective delivery of antifibrotic drugs to rat and human hepatic stellate cells. Hepatology. 1999 May;29(5):1486-93. | | 3. van der Ploeg AT, van der Kraaij AM, Willemsen R, Kroos MA, Loonen MC, Koster JF, Reuser AJ: Rat heart perfusion as model system for enzyme replacement therapy in glycogenosis type II. Pediatr Res. 1990 Oct;28(4):344-7. | | 4. DeRossi C, Bode L, Eklund EA, Zhang F, Davis JA, Westphal V, Wang L, Borowsky AD, Freeze HH: Ablation of mouse phosphomannose isomerase (Mpi) causes mannose 6-phosphate accumulation, toxicity, and embryonic lethality. J Biol Chem. 2006 Mar 3;281(9):5916-27. Epub 2005 Dec 8. | | 5. Yatziv S, Barfi G, Newburg DS: Lysosomal hydrolases in blood-derived macrophages of patients with I-cell disease. J Lab Clin Med. 1986 Oct;108(4):365-8. | | 6. Tiede S, Muschol N, Reutter G, Cantz M, Ullrich K, Braulke T: Missense mutations in N-acetylglucosamine-1-phosphotransferase alpha/beta subunit gene in a patient with mucolipidosis III and a mild clinical phenotype. Am J Med Genet A. 2005 Sep 1;137A(3):235-40. | | 7. Puolakkainen M, Kuo CC, Campbell LA: Chlamydia pneumoniae uses the mannose 6-phosphate/insulin-like growth factor 2 receptor for infection of endothelial cells. Infect Immun. 2005 Aug;73(8):4620-5. | | 8. Holtta-Vuori M, Maatta J, Ullrich O, Kuismanen E, Ikonen E: Mobilization of late-endosomal cholesterol is inhibited by Rab guanine nucleotide dissociation inhibitor. Curr Biol. 2000 Jan 27;10(2):95-8. | | 9. Harper J, Burns JL, Foulstone EJ, Pignatelli M, Zaina S, Hassan AB: Soluble IGF2 receptor rescues Apc(Min/+) intestinal adenoma progression induced by Igf2 loss of imprinting. Cancer Res. 2006 Feb 15;66(4):1940-8. | | 10. Saris JJ, Derkx FH, De Bruin RJ, Dekkers DH, Lamers JM, Saxena PR, Schalekamp MA, Jan Danser AH: High-affinity prorenin binding to cardiac man-6-P/IGF-II receptors precedes proteolytic activation to renin. Am J Physiol Heart Circ Physiol. 2001 Apr;280(4):H1706-15. | | 11. Bird CH, Sun J, Ung K, Karambalis D, Whisstock JC, Trapani JA, Bird PI: Cationic sites on granzyme B contribute to cytotoxicity by promoting its uptake into target cells. Mol Cell Biol. 2005 Sep;25(17):7854-67. | | 12. Adrian JE, Poelstra K, Scherphof GL, Molema G, Meijer DK, Reker-Smit C, Morselt HW, Kamps JA: Interaction of targeted liposomes with primary cultured hepatic stellate cells: Involvement of multiple receptor systems. J Hepatol. 2006 Mar;44(3):560-7. Epub 2005 Oct 19. | | 13. Ullrich K, Basner R, Gieselmann V, Von Figura K: Recognition of human urine alpha-N-acetylglucosaminidase by rat hepatocytes. Involvement of receptors specific for galactose, mannose 6-phosphate and mannose. Biochem J. 1979 May 15;180(2):413-9. | | 14. Davis JA, Wu XH, Wang L, DeRossi C, Westphal V, Wu R, Alton G, Srikrishna G, Freeze HH: Molecular cloning, gene organization, and expression of mouse Mpi encoding phosphomannose isomerase. Glycobiology. 2002 Jul;12(7):435-42. | | 15. Kaplan A, Fischer D, Achord D, Sly W: Phosphohexosyl recognition is a general characteristic of pinocytosis of lysosomal glycosidases by human fibroblasts. J Clin Invest. 1977 Nov;60(5):1088-93. | | 16. Lobel P, Dahms NM, Breitmeyer J, Chirgwin JM, Kornfeld S: Cloning of the bovine 215-kDa cation-independent mannose 6-phosphate receptor. Proc Natl Acad Sci U S A. 1987 Apr;84(8):2233-7. | | 17. Maguchi S, Taniguchi N, Makita A: Elevated activity and increased mannose-6-phosphate in the carbohydrate moiety of cathepsin D from human hepatoma. Cancer Res. 1988 Jan 15;48(2):362-7. | | 18. Sleat DE, Wang Y, Sohar I, Lackland H, Li Y, Li H, Zheng H, Lobel P: Identification and validation of mannose 6-phosphate glycoproteins in human plasma reveal a wide range of lysosomal and non-lysosomal proteins. Mol Cell Proteomics. 2006 Oct;5(10):1942-56. Epub 2006 May 17. | | 19. Sleat DE, Sohar I, Lackland H, Majercak J, Lobel P: Rat brain contains high levels of mannose-6-phosphorylated glycoproteins including lysosomal enzymes and palmitoyl-protein thioesterase, an enzyme implicated in infantile neuronal lipofuscinosis. J Biol Chem. 1996 Aug 9;271(32):19191-8. | | 20. Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. |
|
|---|