| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:27:19 UTC |
|---|
| Update Date | 2016-11-09 01:21:14 UTC |
|---|
| Accession Number | CHEM035059 |
|---|
| Identification |
|---|
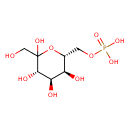
| Common Name | D-Sedoheptulose 7-phosphate |
|---|
| Class | Small Molecule |
|---|
| Description | D-D-d-sedoheptulose 7-phosphate, also known as D-sedoheptulose-7-P, belongs to the class of organic compounds known as hexose phosphates. These are carbohydrate derivatives containing a hexose substituted by one or more phosphate groups. D-D-d-sedoheptulose 7-phosphate is possibly soluble (in water) and an extremely weak basic (essentially neutral) compound (based on its pKa). D-D-d-sedoheptulose 7-phosphate exists in all eukaryotes, ranging from yeast to humans. D-D-d-sedoheptulose 7-phosphate participates in a number of enzymatic reactions, within cattle. In particular, D-Glyceraldehyde 3-phosphate and D-d-sedoheptulose 7-phosphate can be converted into D-ribose 5-phosphate and xylulose 5-phosphate through its interaction with the enzyme transketolase. Furthermore, D-Glyceraldehyde 3-phosphate and D-d-sedoheptulose 7-phosphate can be biosynthesized from D-erythrose 4-phosphate and fructose 6-phosphate; which is mediated by the enzyme transaldolase. Furthermore, D-Glyceraldehyde 3-phosphate and D-d-sedoheptulose 7-phosphate can be biosynthesized from D-erythrose 4-phosphate and fructose 6-phosphate through its interaction with the enzyme transaldolase. Finally, D-Glyceraldehyde 3-phosphate and D-d-sedoheptulose 7-phosphate can be converted into D-ribose 5-phosphate and xylulose 5-phosphate; which is catalyzed by the enzyme transketolase. In cattle, D-d-sedoheptulose 7-phosphate is involved in a couple of metabolic pathways, which include the pentose phosphate pathway and cancer (via the Warburg effect). |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| D-Sedoheptulose 7-phosphoric acid | Generator | | 7-(Dihydrogen phosphate) sedoheptulose | HMDB | | D-Sedoheptulose-7-p | HMDB | | D-Sedoheptulose-7-phosphate | HMDB | | Heptulose-7-phosphate | HMDB | | Sedoheptulose 7-phosphate | HMDB | | Sedoheptulose-7-p | HMDB | | Sedoheptulose-7-phosphate | HMDB |
|
|---|
| Chemical Formula | C7H15O10P |
|---|
| Average Molecular Mass | 290.162 g/mol |
|---|
| Monoisotopic Mass | 290.040 g/mol |
|---|
| CAS Registry Number | 2646-35-7 |
|---|
| IUPAC Name | {[(2R,3S,4R,5S)-3,4,5,6-tetrahydroxy-6-(hydroxymethyl)oxan-2-yl]methoxy}phosphonic acid |
|---|
| Traditional Name | [(2R,3S,4R,5S)-3,4,5,6-tetrahydroxy-6-(hydroxymethyl)oxan-2-yl]methoxyphosphonic acid |
|---|
| SMILES | OCC1(O)O[C@H](COP(O)(O)=O)[C@@H](O)[C@@H](O)[C@@H]1O |
|---|
| InChI Identifier | InChI=1S/C7H15O10P/c8-2-7(12)6(11)5(10)4(9)3(17-7)1-16-18(13,14)15/h3-6,8-12H,1-2H2,(H2,13,14,15)/t3-,4-,5-,6+,7?/m1/s1 |
|---|
| InChI Key | CBIDVWSRUUODHL-QTSLKERKSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as hexose phosphates. These are carbohydrate derivatives containing a hexose substituted by one or more phosphate groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Hexose phosphates |
|---|
| Alternative Parents | |
|---|
| Substituents | - Hexose phosphate
- C-glycosyl compound
- Glycosyl compound
- Monosaccharide phosphate
- Monoalkyl phosphate
- Organic phosphoric acid derivative
- Oxane
- Phosphoric acid ester
- Alkyl phosphate
- Hemiacetal
- Secondary alcohol
- Organoheterocyclic compound
- Oxacycle
- Polyol
- Organic oxide
- Alcohol
- Primary alcohol
- Hydrocarbon derivative
- Aliphatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aliphatic heteromonocyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-05u2-9760000000-23220eac824fb093768e | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (5 TMS) - 70eV, Positive | splash10-000i-4300149000-6fba19ca6cd4460a42b0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0006-1490000000-d647c8b819afa525ac31 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-006y-6940000000-948baa6c0020169263a3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0uk9-6900000000-f2ead32bc69a7827f273 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-009i-8790000000-18c40c04e91399064ede | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9100000000-b41f9c2b1d3edf3ab828 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9000000000-63453353080e28955933 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0001068 |
|---|
| FooDB ID | FDB022407 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | 34485 |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 17216052 |
|---|
| ChEBI ID | 15721 |
|---|
| PubChem Compound ID | 22833559 |
|---|
| Kegg Compound ID | C05382 |
|---|
| YMDB ID | YMDB00196 |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Huck JH, Struys EA, Verhoeven NM, Jakobs C, van der Knaap MS: Profiling of pentose phosphate pathway intermediates in blood spots by tandem mass spectrometry: application to transaldolase deficiency. Clin Chem. 2003 Aug;49(8):1375-80. | | 2. Thornalley PJ, Jahan I, Ng R: Suppression of the accumulation of triosephosphates and increased formation of methylglyoxal in human red blood cells during hyperglycaemia by thiamine in vitro. J Biochem. 2001 Apr;129(4):543-9. | | 3. Nakayama Y, Kinoshita A, Tomita M: Dynamic simulation of red blood cell metabolism and its application to the analysis of a pathological condition. Theor Biol Med Model. 2005 May 9;2:18. | | 4. Makarov SA, Kudriavtseva GV, Kolotilova AI: [Effect of prostaglandins F2 and F2 alpha on the pentosephosate pathway in human blood platelets]. Vopr Med Khim. 1983 Sep-Oct;29(5):27-32. | | 5. Wamelink MM, Struys EA, Huck JH, Roos B, van der Knaap MS, Jakobs C, Verhoeven NM: Quantification of sugar phosphate intermediates of the pentose phosphate pathway by LC-MS/MS: application to two new inherited defects of metabolism. J Chromatogr B Analyt Technol Biomed Life Sci. 2005 Aug 25;823(1):18-25. Epub 2005 Jan 23. |
|
|---|