| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:21:06 UTC |
|---|
| Update Date | 2016-11-09 01:21:13 UTC |
|---|
| Accession Number | CHEM034939 |
|---|
| Identification |
|---|
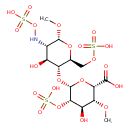
| Common Name | Heparan sulfate |
|---|
| Class | Small Molecule |
|---|
| Description | |
|---|
| Contaminant Sources | - FooDB Chemicals
- STOFF IDENT Compounds
|
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| Heparan sulfuric acid | Generator | | Heparan sulphate | Generator | | Heparan sulphuric acid | Generator | | alpha-Idosane | HMDB | | Heparan N-sulfate | HMDB | | Heparan N-sulphate | HMDB | | Heparatan sulfate | HMDB | | Heparatan sulphate | HMDB | | Heparitin | HMDB | | Heparitin monosulfate | HMDB | | Heparitin monosulphate | HMDB | | Heparitin sulfate | HMDB | | Heparitin sulphate | HMDB | | HHS 5 | HMDB | | N-Acetylheparan sulfate | HMDB | | N-Acetylheparan sulphate | HMDB | | Suleparoid | HMDB | | Tavidan | HMDB | | (2S,3R,4R,5S,6R)-4-Hydroxy-6-{[(2S,3R,4S,5S,6R)-4-hydroxy-6-methoxy-5-[(sulfooxy)amino]-2-[(sulfooxy)methyl]oxan-3-yl]oxy}-3-methoxy-5-(sulfooxy)oxane-2-carboxylate | HMDB | | (2S,3R,4R,5S,6R)-4-Hydroxy-6-{[(2S,3R,4S,5S,6R)-4-hydroxy-6-methoxy-5-[(sulphooxy)amino]-2-[(sulphooxy)methyl]oxan-3-yl]oxy}-3-methoxy-5-(sulphooxy)oxane-2-carboxylate | HMDB | | (2S,3R,4R,5S,6R)-4-Hydroxy-6-{[(2S,3R,4S,5S,6R)-4-hydroxy-6-methoxy-5-[(sulphooxy)amino]-2-[(sulphooxy)methyl]oxan-3-yl]oxy}-3-methoxy-5-(sulphooxy)oxane-2-carboxylic acid | HMDB |
|
|---|
| Chemical Formula | C14H23NO21S3 |
|---|
| Average Molecular Mass | 637.522 g/mol |
|---|
| Monoisotopic Mass | 636.992 g/mol |
|---|
| CAS Registry Number | 9050-30-0 |
|---|
| IUPAC Name | (2S,3R,4R,5S,6R)-4-hydroxy-6-{[(2S,3R,4S,5S,6R)-4-hydroxy-6-methoxy-5-[(sulfooxy)amino]-2-[(sulfooxy)methyl]oxan-3-yl]oxy}-3-methoxy-5-(sulfooxy)oxane-2-carboxylic acid |
|---|
| Traditional Name | (2S,3R,4R,5S,6R)-4-hydroxy-6-{[(2S,3R,4S,5S,6R)-4-hydroxy-6-methoxy-5-[(sulfooxy)amino]-2-[(sulfooxy)methyl]oxan-3-yl]oxy}-3-methoxy-5-(sulfooxy)oxane-2-carboxylic acid |
|---|
| SMILES | Not Available |
|---|
| InChI Identifier | InChI=1S/C14H24NO21S3/c1-29-9-7(17)10(35-38(23,24)25)14(34-11(9)12(18)19)33-8-4(3-31-37(20,21)22)32-13(30-2)5(6(8)16)15-36-39(26,27)28/h4-11,13-15,17H,3H2,1-2H3,(H,18,19)(H,20,21,22)(H,23,24,25)(H,26,27,28)/q-1/p-1/t4-,5-,6-,7+,8-,9+,10-,11-,13+,14+/m0/s1 |
|---|
| InChI Key | WKPUACLQLIIVJJ-RHKLHVFKSA-M |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as disaccharide sulfates. These are disaccharides carrying one or more sulfate group on a sugar unit. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Disaccharide sulfates |
|---|
| Alternative Parents | |
|---|
| Substituents | - Disaccharide sulfate
- 1-o-glucuronide
- O-glucuronide
- Glucuronic acid or derivatives
- Glycosyl compound
- O-glycosyl compound
- Sulfuric acid ester
- Alkyl sulfate
- Sulfate-ester
- Sulfuric acid monoester
- Pyran
- Oxane
- Organic sulfuric acid or derivatives
- Secondary alcohol
- Acetal
- Carboxylic acid derivative
- Carboxylic acid
- Dialkyl ether
- Ether
- Monocarboxylic acid or derivatives
- Oxacycle
- Organoheterocyclic compound
- Organic nitrogen compound
- Organopnictogen compound
- Organic oxide
- Hydrocarbon derivative
- Carbonyl group
- Alcohol
- Organonitrogen compound
- Aliphatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aliphatic heteromonocyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_7) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_7) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_8) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_9) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_10) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_11) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_12) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_13) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_14) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_15) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_16) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_17) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_18) - 70eV, Positive | Not Available | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-0kai-9400000000-404fa85a4342eafffaec | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-001i-9400000000-eec3b32ccab8e708c47b | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-001i-9400000000-ad221bda41e892ec4944 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00dl-0029057000-552f350b2eda79515ae3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0fk9-1079141000-0b28aff1bbd9804b9bea | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-05fr-1394000000-5fa03687957fac5d0d21 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-00kr-2923734000-0b76c8fcf07de6f623f2 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-014i-4903331000-2468d6710f7d484c4bc2 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-00xr-7901000000-aa930e030c48c4c24016 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0000019000-61194d75acd7b76eeb63 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000b-9380064000-507abf4005bb44c55d09 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0002-9232401000-8b2d59e928ea5f7d2bf8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0006-0000009000-37a3f7caf057b5f5103f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0006-3061394000-8c7732b97f391e56ce66 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0pb9-3291100000-baca1b5774273b60adc0 | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 2D NMR | [1H,13C] 2D NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0000693 |
|---|
| FooDB ID | FDB022184 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Heparan sulfate |
|---|
| Chemspider ID | Not Available |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 53477715 |
|---|
| Kegg Compound ID | C00925 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Meier, Bernd Horst; Meier, Nele. Manufacture and use of modified adducts of polysaccharides and chitosan or chitin and a process to improve the preparation of polysaccharide-drug conjugates. PCT Int. Appl. (2007), 67pp. CODEN: PIXXD2 WO 2007122269 A1 20071101 CAN 147:487869 AN 2007:1237349 | | 2. Inoue H, Otsu K, Yoneda M, Kimata K, Suzuki S, Nakanishi Y: Glycosaminoglycan sulfotransferases in human and animal sera. J Biol Chem. 1986 Apr 5;261(10):4460-9. | | 3. Nader HB, Lopes CC, Rocha HA, Santos EA, Dietrich CP: Heparins and heparinoids: occurrence, structure and mechanism of antithrombotic and hemorrhagic activities. Curr Pharm Des. 2004;10(9):951-66. | | 4. Henriquez JP, Casar JC, Fuentealba L, Carey DJ, Brandan E: Extracellular matrix histone H1 binds to perlecan, is present in regenerating skeletal muscle and stimulates myoblast proliferation. J Cell Sci. 2002 May 15;115(Pt 10):2041-51. | | 5. Hjelm Cluff A, Malmstrom A, Tingaker B, David G, Ekman-Ordeberg G: Normal labor associated with changes in uterine heparan sulfate proteoglycan expression and localization. Acta Obstet Gynecol Scand. 2005 Mar;84(3):217-24. | | 6. Yu WH, Yu S, Meng Q, Brew K, Woessner JF Jr: TIMP-3 binds to sulfated glycosaminoglycans of the extracellular matrix. J Biol Chem. 2000 Oct 6;275(40):31226-32. | | 7. Subramanian G, LeBlanc RA, Wardley RC, Fuller AO: Defective entry of herpes simplex virus types 1 and 2 into porcine cells and lack of infection in infant pigs indicate species tropism. J Gen Virol. 1995 Sep;76 ( Pt 9):2375-9. | | 8. Jenniskens GJ, Veerkamp JH, van Kuppevelt TH: Heparan sulfates in skeletal muscle development and physiology. J Cell Physiol. 2006 Feb;206(2):283-94. | | 9. Maeda T, Alexander CM, Friedl A: Induction of syndecan-1 expression in stromal fibroblasts promotes proliferation of human breast cancer cells. Cancer Res. 2004 Jan 15;64(2):612-21. | | 10. Sher I, Zisman-Rozen S, Eliahu L, Whitelock JM, Maas-Szabowski N, Yamada Y, Breitkreutz D, Fusenig NE, Arikawa-Hirasawa E, Iozzo RV, Bergman R, Ron D: Targeting perlecan in human keratinocytes reveals novel roles for perlecan in epidermal formation. J Biol Chem. 2006 Feb 24;281(8):5178-87. Epub 2005 Nov 2. | | 11. Savas PS, Hemsley KM, Hopwood JJ: Intracerebral injection of sulfamidase delays neuropathology in murine MPS-IIIA. Mol Genet Metab. 2004 Aug;82(4):273-85. | | 12. Chernousov MA, Rothblum K, Tyler WA, Stahl RC, Carey DJ: Schwann cells synthesize type V collagen that contains a novel alpha 4 chain. Molecular cloning, biochemical characterization, and high affinity heparin binding of alpha 4(V) collagen. J Biol Chem. 2000 Sep 8;275(36):28208-15. | | 13. Chikama S, Iida S, Inoue M, Kawagoe N, Tomiyasu K, Matsuoka K, Noda S, Takazono I: Role of heparan sulfate proteoglycan (syndecan-1) on the renal epithelial cells during calcium oxalate monohydrate crystal attachment. Kurume Med J. 2002;49(4):201-10. | | 14. Kosir MA, Foley-Loudon PA, Finkenauer R, Tennenberg SD: Multiple heparanases are expressed in polymorphonuclear cells. J Surg Res. 2002 Mar;103(1):100-8. | | 15. Myette JR, Shriver Z, Liu J, Venkataraman G, Rosenberg R, Sasisekharan R: Expression in Escherichia coli, purification and kinetic characterization of human heparan sulfate 3-O-sulfotransferase-1. Biochem Biophys Res Commun. 2002 Feb 1;290(4):1206-13. | | 16. Barnett MW, Fisher CE, Perona-Wright G, Davies JA: Signalling by glial cell line-derived neurotrophic factor (GDNF) requires heparan sulphate glycosaminoglycan. J Cell Sci. 2002 Dec 1;115(Pt 23):4495-503. | | 17. Wasserman L, Abramovici A, Shlesinger H, Goldman JA, Allalouf D: Histochemical localization of acidic glycosaminoglycans in normal human placentae. Placenta. 1983 Jan-Apr;4(1):101-8. | | 18. Haimov-Kochman R, Friedmann Y, Prus D, Goldman-Wohl DS, Greenfield C, Anteby EY, Aviv A, Vlodavsky I, Yagel S: Localization of heparanase in normal and pathological human placenta. Mol Hum Reprod. 2002 Jun;8(6):566-73. | | 19. Dempsey LA, Plummer TB, Coombes SL, Platt JL: Heparanase expression in invasive trophoblasts and acute vascular damage. Glycobiology. 2000 May;10(5):467-75. | | 20. Leu SJ, Chen N, Chen CC, Todorovic V, Bai T, Juric V, Liu Y, Yan G, Lam SC, Lau LF: Targeted mutagenesis of the angiogenic protein CCN1 (CYR61). Selective inactivation of integrin alpha6beta1-heparan sulfate proteoglycan coreceptor-mediated cellular functions. J Biol Chem. 2004 Oct 15;279(42):44177-87. Epub 2004 Aug 17. | | 21. Lensen JF, Rops AL, Wijnhoven TJ, Hafmans T, Feitz WF, Oosterwijk E, Banas B, Bindels RJ, van den Heuvel LP, van der Vlag J, Berden JH, van Kuppevelt TH: Localization and functional characterization of glycosaminoglycan domains in the normal human kidney as revealed by phage display-derived single chain antibodies. J Am Soc Nephrol. 2005 May;16(5):1279-88. Epub 2005 Mar 23. |
|
|---|