| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:13:44 UTC |
|---|
| Update Date | 2016-11-09 01:21:11 UTC |
|---|
| Accession Number | CHEM034795 |
|---|
| Identification |
|---|
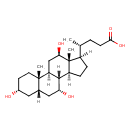
| Common Name | 3a,7a,12b-Trihydroxy-5b-cholanoic acid |
|---|
| Class | Small Molecule |
|---|
| Description | |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 3alpha,7alpha,12beta-Trihydroxy-5beta-cholanate | ChEBI | | Lagocholic acid | ChEBI | | 3a,7a,12b-Trihydroxy-5b-cholanate | Generator | | 3a,7a,12b-Trihydroxy-5b-cholanic acid | Generator | | 3alpha,7alpha,12beta-Trihydroxy-5beta-cholanic acid | Generator | | 3α,7α,12β-trihydroxy-5β-cholanate | Generator | | 3α,7α,12β-trihydroxy-5β-cholanic acid | Generator | | 3a,7a,12b-Trihydroxy-5b-cholanoate | Generator, HMDB | | Lagocholate | Generator | | 12b-Hydroxyisocholate | HMDB | | 12b-Hydroxyisocholic acid | HMDB | | (4R)-4-[(1S,2S,5R,7S,9R,10R,11S,14R,15R,16R)-5,9,16-Trihydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]pentanoate | Generator, HMDB | | (3alpha,5beta,7alpha,12beta)-3,7,12-Trihydroxycholan-24-oic acid | HMDB | | (3α,5β,7α,12β)-3,7,12-Trihydroxycholan-24-oic acid | HMDB | | 12beta-Hydroxyisocholic acid | HMDB | | 12β-Hydroxyisocholic acid | HMDB | | 3alpha,7alpha,12beta-Trihydroxy-5beta-cholanoic acid | HMDB | | 3α,7α,12β-Trihydroxy-5β-cholanoic acid | HMDB |
|
|---|
| Chemical Formula | C24H40O5 |
|---|
| Average Molecular Mass | 408.571 g/mol |
|---|
| Monoisotopic Mass | 408.288 g/mol |
|---|
| CAS Registry Number | 71883-64-2 |
|---|
| IUPAC Name | (4R)-4-[(1S,2S,5R,7S,9R,10R,11S,14R,15R,16R)-5,9,16-trihydroxy-2,15-dimethyltetracyclo[8.7.0.0^{2,7}.0^{11,15}]heptadecan-14-yl]pentanoic acid |
|---|
| Traditional Name | (4R)-4-[(1S,2S,5R,7S,9R,10R,11S,14R,15R,16R)-5,9,16-trihydroxy-2,15-dimethyltetracyclo[8.7.0.0^{2,7}.0^{11,15}]heptadecan-14-yl]pentanoic acid |
|---|
| SMILES | [H][C@@]12CC[C@H]([C@H](C)CCC(O)=O)[C@@]1(C)[C@H](O)C[C@@]1([H])[C@@]2([H])[C@H](O)C[C@]2([H])C[C@H](O)CC[C@]12C |
|---|
| InChI Identifier | InChI=1S/C24H40O5/c1-13(4-7-21(28)29)16-5-6-17-22-18(12-20(27)24(16,17)3)23(2)9-8-15(25)10-14(23)11-19(22)26/h13-20,22,25-27H,4-12H2,1-3H3,(H,28,29)/t13-,14+,15-,16-,17+,18+,19-,20-,22+,23+,24-/m1/s1 |
|---|
| InChI Key | BHQCQFFYRZLCQQ-KRHHAYMPSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as trihydroxy bile acids, alcohols and derivatives. These are prenol lipids structurally characterized by a bile acid or alcohol which bears three hydroxyl groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Steroids and steroid derivatives |
|---|
| Sub Class | Bile acids, alcohols and derivatives |
|---|
| Direct Parent | Trihydroxy bile acids, alcohols and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Trihydroxy bile acid, alcohol, or derivatives
- 3-hydroxysteroid
- 12-hydroxysteroid
- 7-hydroxysteroid
- 3-alpha-hydroxysteroid
- Hydroxysteroid
- Cyclic alcohol
- Secondary alcohol
- Carboxylic acid derivative
- Carboxylic acid
- Polyol
- Monocarboxylic acid or derivatives
- Organic oxide
- Alcohol
- Organic oxygen compound
- Hydrocarbon derivative
- Carbonyl group
- Organooxygen compound
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-03ec-0439000000-abd2a5c162e23ea0d78f | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (4 TMS) - 70eV, Positive | splash10-001i-1100049000-86faaffe1dc0eadf9e75 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_3) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00dl-0009000000-f5b70cbc59d5ea066854 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00dl-0009000000-b06963261afb6fdb494b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-02c2-2109000000-87477f9c822aeda6a8e0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4r-0009700000-8dc05bd9542a54f1f91b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-052r-0009200000-9a24af3f58b590ac86cd | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4l-9007000000-c0f6af27285559bd789e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0001900000-a764d2a5e3d1918ae5fa | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4r-0007900000-d1ed92b278bc96ae6ebd | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-1019600000-c907470476ed433e714e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0abc-0019500000-627bd12690a4dc559a07 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-052r-4379100000-2566436d8a667838d682 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4j-8930000000-8acc72953d3f45da3f73 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0000312 |
|---|
| FooDB ID | FDB021939 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | 5301 |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 4446957 |
|---|
| ChEBI ID | 36240 |
|---|
| PubChem Compound ID | 5283869 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | ECMDB24033 |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Batta, Ashok K.; Aggarwal, Suresht K.; Salen, Gerald; Shefer, Sarah. Selective reduction of oxo bile acids: synthesis of 3b-, 7b-, and 12b-hydroxy bile acids. Journal of Lipid Research (1991), 32(6), 977-83. | | 2. Borgstrom B, Krabisch L, Lindstrom M: Metabolism of cholate, 7 beta-hydroxy- and 12 beta-hydroxy-isocholates in the rat. Scand J Clin Lab Invest. 1986 Apr;46(2):177-82. | | 3. Yamaga N, Ikebuchi J, Kohara H, Ogura Y, Yamada K: Analysis of bile acids in urine specimens from healthy humans: determination of several bile acids with beta-hydroxyl and carbonyl groups. J Biochem. 1996 Apr;119(4):725-30. | | 4. St-Pierre MV, Kullak-Ublick GA, Hagenbuch B, Meier PJ: Transport of bile acids in hepatic and non-hepatic tissues. J Exp Biol. 2001 May;204(Pt 10):1673-86. | | 5. Claudel T, Staels B, Kuipers F: The Farnesoid X receptor: a molecular link between bile acid and lipid and glucose metabolism. Arterioscler Thromb Vasc Biol. 2005 Oct;25(10):2020-30. Epub 2005 Jul 21. | | 6. Chiang JY: Bile acid regulation of hepatic physiology: III. Bile acids and nuclear receptors. Am J Physiol Gastrointest Liver Physiol. 2003 Mar;284(3):G349-56. | | 7. Davis RA, Miyake JH, Hui TY, Spann NJ: Regulation of cholesterol-7alpha-hydroxylase: BAREly missing a SHP. J Lipid Res. 2002 Apr;43(4):533-43. | | 8. https://www.ncbi.nlm.nih.gov/pubmed/?term=11316487 | | 9. https://www.ncbi.nlm.nih.gov/pubmed/?term=16037564 |
|
|---|