| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 05:13:08 UTC |
|---|
| Update Date | 2016-11-09 01:21:11 UTC |
|---|
| Accession Number | CHEM034785 |
|---|
| Identification |
|---|
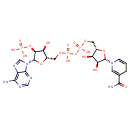
| Common Name | NADPH |
|---|
| Class | Small Molecule |
|---|
| Description | |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 2'-(Dihydrogen phosphate) 5'-(trihydrogen pyrophosphate) adenosine 5'-ester with 1,4-dihydro-1-b-D-ribofuranosylnicotinamide | HMDB | | 2'-(Dihydrogen phosphate) 5'-(trihydrogen pyrophosphate) adenosine 5'-ester with 1,4-dihydro-1-beta-delta-ribofuranosylnicotinamide | HMDB | | Adenosine 5'-(trihydrogen diphosphate) 2'-(dihydrogen phosphate) p'-5'-ester with 1,4-dihydro-1-beta-D-ribofuranosyl-3-pyridinecarboxamide | HMDB | | Adenosine 5'-(trihydrogen diphosphate) 2'-(dihydrogen phosphate) p'-5'-ester with 1,4-dihydro-1-beta-delta-ribofuranosyl-3-pyridinecarboxamide | HMDB | | b-NADPH | HMDB | | b-Nicotinamide-adenine-dinucleotide-phosphorate | HMDB | | b-Nicotinamide-adenine-dinucleotide-phosphoric acid | HMDB | | beta-NADPH | HMDB | | beta-Nicotinamide-adenine-dinucleotide-phosphorate | HMDB | | beta-Nicotinamide-adenine-dinucleotide-phosphoric acid | HMDB | | Dihydrocodehydrogenase II | HMDB | | Dihydronicotinamide adenine dinucleotide phosphate | HMDB | | Dihydronicotinamide adenine dinucleotide-p | HMDB | | Dihydrotriphosphopyridine nucleotide reduced | HMDB | | NADP-reduced | HMDB | | Nicotinamide adenine dinucleotide phosphate - reduced | HMDB | | Nicotinamide-adenine-dinucleotide-phosphorate | HMDB | | Nicotinamide-adenine-dinucleotide-phosphoric acid | HMDB | | Reduced codehydrase II | HMDB | | Reduced coenzyme II | HMDB | | Reduced cozymase II | HMDB | | Reduced triphosphopyridine nucleotide | HMDB | | Triphosphopyridine nucleotide reduced | HMDB |
|
|---|
| Chemical Formula | C21H30N7O17P3 |
|---|
| Average Molecular Mass | 745.421 g/mol |
|---|
| Monoisotopic Mass | 745.091 g/mol |
|---|
| CAS Registry Number | 53-57-6 |
|---|
| IUPAC Name | {[(2S,3S,4S,5S)-2-(6-amino-9H-purin-9-yl)-5-[({[({[(2S,3R,4S,5S)-5-(3-carbamoyl-1,4-dihydropyridin-1-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}(hydroxy)phosphoryl)oxy](hydroxy)phosphoryl}oxy)methyl]-4-hydroxyoxolan-3-yl]oxy}phosphonic acid |
|---|
| Traditional Name | [(2S,3S,4S,5S)-2-(6-aminopurin-9-yl)-5-{[({[(2S,3R,4S,5S)-5-(3-carbamoyl-4H-pyridin-1-yl)-3,4-dihydroxyoxolan-2-yl]methoxy(hydroxy)phosphoryl}oxy(hydroxy)phosphoryl)oxy]methyl}-4-hydroxyoxolan-3-yl]oxyphosphonic acid |
|---|
| SMILES | NC(=O)C1=CN(C=CC1)[C@H]1O[C@@H](COP(O)(=O)OP(O)(=O)OC[C@@H]2O[C@@H]([C@@H](OP(O)(O)=O)[C@H]2O)N2C=NC3=C(N)N=CN=C23)[C@H](O)[C@@H]1O |
|---|
| InChI Identifier | InChI=1S/C21H30N7O17P3/c22-17-12-19(25-7-24-17)28(8-26-12)21-16(44-46(33,34)35)14(30)11(43-21)6-41-48(38,39)45-47(36,37)40-5-10-13(29)15(31)20(42-10)27-3-1-2-9(4-27)18(23)32/h1,3-4,7-8,10-11,13-16,20-21,29-31H,2,5-6H2,(H2,23,32)(H,36,37)(H,38,39)(H2,22,24,25)(H2,33,34,35)/t10-,11-,13-,14-,15-,16-,20-,21-/m0/s1 |
|---|
| InChI Key | ACFIXJIJDZMPPO-NCHANQSKSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as (5'->5')-dinucleotides. These are dinucleotides where the two bases are connected via a (5'->5')-phosphodiester linkage. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Nucleosides, nucleotides, and analogues |
|---|
| Class | (5'->5')-dinucleotides |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | (5'->5')-dinucleotides |
|---|
| Alternative Parents | |
|---|
| Substituents | - (5'->5')-dinucleotide
- Purine nucleotide sugar
- Purine ribonucleoside 2',5'-bisphosphate
- Purine ribonucleoside bisphosphate
- Purine ribonucleoside diphosphate
- Nicotinamide-nucleotide
- Pentose phosphate
- Pentose-5-phosphate
- N-glycosyl compound
- Glycosyl compound
- Monosaccharide phosphate
- 6-aminopurine
- N-substituted nicotinamide
- Organic pyrophosphate
- Imidazopyrimidine
- Purine
- Monoalkyl phosphate
- Dihydropyridine
- Aminopyrimidine
- Organic phosphoric acid derivative
- Pyrimidine
- N-substituted imidazole
- Hydropyridine
- Alkyl phosphate
- Phosphoric acid ester
- Imidolactam
- Monosaccharide
- Vinylogous amide
- Heteroaromatic compound
- Tetrahydrofuran
- Azole
- Imidazole
- Amino acid or derivatives
- Carboxamide group
- Secondary alcohol
- Primary carboxylic acid amide
- Oxacycle
- Azacycle
- Organoheterocyclic compound
- Carboxylic acid derivative
- Enamine
- Hydrocarbon derivative
- Organic oxygen compound
- Amine
- Organopnictogen compound
- Carbonyl group
- Organic oxide
- Primary amine
- Alcohol
- Organooxygen compound
- Organonitrogen compound
- Organic nitrogen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-004j-9421780200-89062ec4778392672634 | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-02tj-1838493720-b38b37650e8976905bb0 | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-0a4r-0841393100-6e4cbc8ac576e3e8b195 | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-0550-2694875730-0dc522520957880672cb | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0921104300-c1952197674952109d2c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0901000000-31adeef6c90e9070dfa4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-0920000000-e29dcab04f59f9c7189c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0036-6900111800-ab3c73d03c4430e1b644 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-003r-4901100000-96b551ade48c419935ed | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-057i-5900000000-0898019fd8b891d5245e | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 2D NMR | [1H,1H] 2D NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0000221 |
|---|
| FooDB ID | FDB021909 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | 33486 |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | 3691 |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Nicotinamide adenine dinucleotide phosphate |
|---|
| Chemspider ID | 17215925 |
|---|
| ChEBI ID | 16474 |
|---|
| PubChem Compound ID | 22833512 |
|---|
| Kegg Compound ID | C00005 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Seelbach, Karsten; Riebel, Bettina; Hummel, Werner; Kula, Maria-Regina; Tishkov, Vladimir I.; Egorov, Alexey M.; Wandrey, Christian; Kragl, Udo. A novel, efficient regenerating method of NADPH using a new formate dehydrogenase. Tetrahedron Letters (1996), 37(9), 1377-80. | | 2. Shindo Y, Witt E, Han D, Epstein W, Packer L: Enzymic and non-enzymic antioxidants in epidermis and dermis of human skin. J Invest Dermatol. 1994 Jan;102(1):122-4. | | 3. Iwata H, Tezuka Y, Kadota S, Hiratsuka A, Watabe T: Mechanism-based inactivation of human liver microsomal CYP3A4 by rutaecarpine and limonin from Evodia fruit extract. Drug Metab Pharmacokinet. 2005 Feb;20(1):34-45. | | 4. Birkmayer GJ, Birkmayer W: Stimulation of endogenous L-dopa biosynthesis--a new principle for the therapy of Parkinson's disease. The clinical effect of nicotinamide adenine dinucleotide (NADH) and nicotinamide adenine dinucleotidephosphate (NADPH). Acta Neurol Scand Suppl. 1989;126:183-7. | | 5. Lee AJ, Zhu BT: NADPH-dependent formation of polar and nonpolar estrogen metabolites following incubations of 17 beta-estradiol with human liver microsomes. Drug Metab Dispos. 2004 Aug;32(8):876-83. | | 6. Kochansky CJ, Xia YQ, Wang S, Cato B, Creighton M, Vincent SH, Franklin RB, Reed JR: Species differences in the elimination of a peroxisome proliferator-activated receptor agonist highlighted by oxidative metabolism of its acyl glucuronide. Drug Metab Dispos. 2005 Dec;33(12):1894-904. Epub 2005 Sep 23. | | 7. Nakayama Y, Kinoshita A, Tomita M: Dynamic simulation of red blood cell metabolism and its application to the analysis of a pathological condition. Theor Biol Med Model. 2005 May 9;2:18. | | 8. Karanam BV, Hop CE, Liu DQ, Wallace M, Dean D, Satoh H, Komuro M, Awano K, Vincent SH: In vitro metabolism of MK-0767 [(+/-)-5-[(2,4-dioxothiazolidin-5-yl)methyl]-2-methoxy-N-[[(4-trifluoromethyl) phenyl]methyl]benzamide], a peroxisome proliferator-activated receptor alpha/gamma agonist. I. Role of cytochrome P450, methyltransferases, flavin monooxygenases, and esterases. Drug Metab Dispos. 2004 Sep;32(9):1015-22. | | 9. Soglia JR, Contillo LG, Kalgutkar AS, Zhao S, Hop CE, Boyd JG, Cole MJ: A semiquantitative method for the determination of reactive metabolite conjugate levels in vitro utilizing liquid chromatography-tandem mass spectrometry and novel quaternary ammonium glutathione analogues. Chem Res Toxicol. 2006 Mar;19(3):480-90. | | 10. Afanas'ev IB, Suslova TB, Cheremisina ZP, Abramova NE, Korkina LG: Study of antioxidant properties of metal aspartates. Analyst. 1995 Mar;120(3):859-62. | | 11. Lin CC, Wong BK, Burgey CS, Gibson CR, Singh R: In vitro metabolism of a thrombin inhibitor and quantitation of metabolically generated cyanide. J Pharm Biomed Anal. 2005 Oct 4;39(5):1014-20. Epub 2005 Jul 14. | | 12. Conley AJ, Pattison JC, Bird IM: Variations in adrenal androgen production among (nonhuman) primates. Semin Reprod Med. 2004 Nov;22(4):311-26. |
|
|---|