| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 04:18:59 UTC |
|---|
| Update Date | 2016-11-09 01:20:58 UTC |
|---|
| Accession Number | CHEM033631 |
|---|
| Identification |
|---|
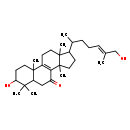
| Common Name | Lucidadiol |
|---|
| Class | Small Molecule |
|---|
| Description | Lucidadiol is found in mushrooms. Lucidadiol is isolated from Ganoderma lucidum (reishi). |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 3,26-Dihydroxylanost-8,24-dien-7-one | HMDB |
|
|---|
| Chemical Formula | C30H48O3 |
|---|
| Average Molecular Mass | 456.700 g/mol |
|---|
| Monoisotopic Mass | 456.360 g/mol |
|---|
| CAS Registry Number | 252351-95-4 |
|---|
| IUPAC Name | 5-hydroxy-14-[(5E)-7-hydroxy-6-methylhept-5-en-2-yl]-2,6,6,11,15-pentamethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadec-1(10)-en-9-one |
|---|
| Traditional Name | 5-hydroxy-14-[(5E)-7-hydroxy-6-methylhept-5-en-2-yl]-2,6,6,11,15-pentamethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadec-1(10)-en-9-one |
|---|
| SMILES | CC(CC\C=C(/C)CO)C1CCC2(C)C3=C(CCC12C)C1(C)CCC(O)C(C)(C)C1CC3=O |
|---|
| InChI Identifier | InChI=1S/C30H48O3/c1-19(18-31)9-8-10-20(2)21-11-16-30(7)26-22(12-15-29(21,30)6)28(5)14-13-25(33)27(3,4)24(28)17-23(26)32/h9,20-21,24-25,31,33H,8,10-18H2,1-7H3/b19-9+ |
|---|
| InChI Key | AZPOACUDFJKUHJ-DJKKODMXSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as triterpenoids. These are terpene molecules containing six isoprene units. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Triterpenoids |
|---|
| Direct Parent | Triterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Triterpenoid
- 26-hydroxysteroid
- Dihydroxy bile acid, alcohol, or derivatives
- Hydroxy bile acid, alcohol, or derivatives
- Bile acid, alcohol, or derivatives
- 3-hydroxysteroid
- Hydroxysteroid
- Oxosteroid
- 7-oxosteroid
- Steroid
- Fatty alcohol
- Cyclohexenone
- Fatty acyl
- Cyclic alcohol
- Ketone
- Secondary alcohol
- Primary alcohol
- Organooxygen compound
- Alcohol
- Hydrocarbon derivative
- Organic oxide
- Carbonyl group
- Organic oxygen compound
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-056u-2006900000-1553271d7f97f7d66372 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-0079-1302490000-39da5f75059c25330ed0 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-052r-0001900000-2ed0a93dcb9c26b3b072 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00dr-1005900000-eb7b7320f5862d3d71fa | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0670-5359800000-6126514566c6bbd99e68 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0000900000-282f558239129ba26104 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4r-0000900000-eae82aa6c173d6619473 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-2003900000-c5fee87caf1adbecd2b2 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4j-9322400000-58cbebb7616d5706c1c6 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4j-9105000000-7fe5433e48732ed1b51c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-056r-9437000000-e59d48484add279531a0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a70-0000900000-b19ae51dfb84827c70f1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a6r-0000900000-21986685affb3d9d212a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0pk9-1002900000-8990a69e270834ee1432 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0040455 |
|---|
| FooDB ID | FDB020208 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Lucidadiol |
|---|
| Chemspider ID | 11596029 |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 22723638 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|