| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 04:18:58 UTC |
|---|
| Update Date | 2016-11-09 01:20:58 UTC |
|---|
| Accession Number | CHEM033630 |
|---|
| Identification |
|---|
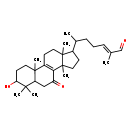
| Common Name | Lucidal |
|---|
| Class | Small Molecule |
|---|
| Description | Lucidal is found in mushrooms. Lucidal is isolated from Ganoderma lucidum (reishi). |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| (+)-Lucialdehyde C | HMDB | | 3-Hydroxy-7-oxolanosta-8,24-dien-26-al | HMDB | | Lucialdehyde C | HMDB, MeSH |
|
|---|
| Chemical Formula | C30H46O3 |
|---|
| Average Molecular Mass | 454.684 g/mol |
|---|
| Monoisotopic Mass | 454.345 g/mol |
|---|
| CAS Registry Number | 252351-96-5 |
|---|
| IUPAC Name | (2E)-6-{5-hydroxy-2,6,6,11,15-pentamethyl-9-oxotetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadec-1(10)-en-14-yl}-2-methylhept-2-enal |
|---|
| Traditional Name | (2E)-6-{5-hydroxy-2,6,6,11,15-pentamethyl-9-oxotetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadec-1(10)-en-14-yl}-2-methylhept-2-enal |
|---|
| SMILES | CC(CC\C=C(/C)C=O)C1CCC2(C)C3=C(CCC12C)C1(C)CCC(O)C(C)(C)C1CC3=O |
|---|
| InChI Identifier | InChI=1S/C30H46O3/c1-19(18-31)9-8-10-20(2)21-11-16-30(7)26-22(12-15-29(21,30)6)28(5)14-13-25(33)27(3,4)24(28)17-23(26)32/h9,18,20-21,24-25,33H,8,10-17H2,1-7H3/b19-9+ |
|---|
| InChI Key | PIOYBULRRJNPSG-DJKKODMXSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as triterpenoids. These are terpene molecules containing six isoprene units. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Triterpenoids |
|---|
| Direct Parent | Triterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Triterpenoid
- 27-oxosteroid
- 26-oxosteroid
- Hydroxy bile acid, alcohol, or derivatives
- Monohydroxy bile acid, alcohol, or derivatives
- Bile acid, alcohol, or derivatives
- 3-hydroxysteroid
- Hydroxysteroid
- 7-oxosteroid
- Oxosteroid
- Steroid
- Cyclohexenone
- Enal
- Cyclic alcohol
- Alpha,beta-unsaturated aldehyde
- Ketone
- Secondary alcohol
- Organooxygen compound
- Organic oxygen compound
- Hydrocarbon derivative
- Carbonyl group
- Organic oxide
- Aldehyde
- Alcohol
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0fbi-0016900000-c8b61a8df35c8496d65f | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-03di-4124890000-06504be1be6e7e015cff | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-052r-1001900000-9b3a2f99c172b1bd8e10 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-05r9-2016900000-afa718760219fd9776a1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-066r-4276900000-58c9d585c024ced18138 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0000900000-0fc02100c44ed9c35378 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0zi9-0000900000-d7d45687d36dc9b3126e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-2002900000-0d4a78310a8a20b2db77 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-052b-9113300000-23146fc8f459421f2598 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-052b-9005100000-a9e5f07f106aa0528169 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-9226000000-8d8d3d73ddc9e633794c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0f79-0000900000-ad5de3c829479306707f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0fi0-0000900000-405b7b6dd26378f1efdc | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0l3s-3004900000-a985144e711018b2ede6 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0040454 |
|---|
| FooDB ID | FDB020207 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00033125 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 8542161 |
|---|
| ChEBI ID | 66594 |
|---|
| PubChem Compound ID | 131752822 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|