| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 04:10:55 UTC |
|---|
| Update Date | 2016-11-09 01:20:55 UTC |
|---|
| Accession Number | CHEM033455 |
|---|
| Identification |
|---|
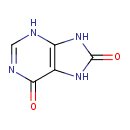
| Common Name | 6,8-Purinediol |
|---|
| Class | Small Molecule |
|---|
| Description | |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 6,8-Purinediol | HMDB | | 7,9-Dihydro-1H-purine-6,8-dione | HMDB | | 7,9-Dihydro-8H-purin-8-one | HMDB | | 8-Oxohypoxanthine | HMDB | | Purine-6,8(1H,9H)-dione | HMDB | | Purine-6,8-diol | HMDB | | Purine-6,8-dione | HMDB |
|
|---|
| Chemical Formula | C25H20N20O10 |
|---|
| Average Molecular Mass | 760.554 g/mol |
|---|
| Monoisotopic Mass | 760.167 g/mol |
|---|
| CAS Registry Number | 13231-00-0 |
|---|
| IUPAC Name | 6,7,8,9-tetrahydro-3H-purine-6,8-dione |
|---|
| Traditional Name | 7,9-dihydro-3H-purine-6,8-dione |
|---|
| SMILES | OC1=NC2=C(N1)NC=NC2=O.OC1=NC2=C(N1)N=CN=C2O.OC1=NC2=C(O)N=CNC2=N1.O=C1NC2=C(N1)C(=O)N=CN2.O=C1NC2=C(N1)C(=O)NC=N2 |
|---|
| InChI Identifier | InChI=1S/5C5H4N4O2/c5*10-4-2-3(6-1-7-4)9-5(11)8-2/h5*1H,(H3,6,7,8,9,10,11) |
|---|
| InChI Key | ZHWYDMYXJSUNAJ-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as hypoxanthines. Hypoxanthines are compounds containing the purine derivative 1H-purin-6(9H)-one. Purine is a bicyclic aromatic compound made up of a pyrimidine ring fused to an imidazole ring. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Imidazopyrimidines |
|---|
| Sub Class | Purines and purine derivatives |
|---|
| Direct Parent | Hypoxanthines |
|---|
| Alternative Parents | |
|---|
| Substituents | - 6-oxopurine
- Hypoxanthine
- Pyrimidone
- Pyrimidine
- Azole
- Imidazole
- Heteroaromatic compound
- Vinylogous amide
- Urea
- Azacycle
- Hydrocarbon derivative
- Organooxygen compound
- Organonitrogen compound
- Organic oxide
- Organopnictogen compound
- Organic oxygen compound
- Organic nitrogen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0zi1-6900000000-b597900216a7c18ff4ec | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0900000000-73408e2a019cf6145389 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-0900000000-4ec7e27bd75758d5243d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000x-9400000000-4b36792f2a9b31c0e39f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0900000000-0c74ea3af474d10b7ea8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0udi-0900000000-30080f0e6c244018af86 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-053u-9600000000-23699c571e13d9a8c982 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0900000000-ac450658003af6c0a109 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-0900000000-00ec07b7bda30887c327 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-05ai-9200000000-f8c6511273a5bfd31f74 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0900000000-757517510c89a3191171 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0fk9-2900000000-b1f56627b7499b83ee76 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9000000000-5b696d1d5422470c88ea | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0001182 |
|---|
| FooDB ID | FDB112171 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 75117 |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 83252 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Ohtsuka, Yozo; Sugimoto, Kikuo. Oxazolopyrimidines. III. Formation of 6,8-dihydroxypurine by the oxidation of 7-aminooxazolo[5,4-d]pyrimidine with hydrogen peroxide-acetic acid. Bulletin of the Chemical Society of Japan (1970), 43(7), 2281. | | 2. Ohtsuka, Yozo; Sugimoto, Kikuo. Oxazolopyrimidines. III. Formation of 6,8-dihydroxypurine by the oxidation of 7-aminooxazolo[5,4-d]pyrimidine with hydrogen peroxide-acetic acid. Bulletin of the Chemical Society of Japan (1970), 43(7), 2281. | | 3. Davies RJ: Complexes of poly(adenylic acid) with complementary monomers. Eur J Biochem. 1976 Jan 2;61(1):225-36. | | 4. TRICERRI S, BRASCHI A: [Quantitative spectrophotometric determination of 6-hydroxypurine in the presence of 2, 6-dihydroxypurine]. Farmaco Sci. 1957;12(1):28-33. | | 5. Smith M, Sullivan C: Germination of Clostridium cylindrosporum Spores on Medium Containing Uric Acid. Appl Environ Microbiol. 1989 Jun;55(6):1380-5. | | 6. Montero-Moran GM, Li M, Rendon-Huerta E, Jourdan F, Lowe DJ, Stumpff-Kane AW, Feig M, Scazzocchio C, Hausinger RP: Purification and characterization of the FeII- and alpha-ketoglutarate-dependent xanthine hydroxylase from Aspergillus nidulans. Biochemistry. 2007 May 8;46(18):5293-304. Epub 2007 Apr 13. | | 7. Salas JA, Johnstone K, Ellar DJ: Role of uricase in the triggering of germination of Bacillus fastidiosus spores. Biochem J. 1985 Jul 1;229(1):241-9. | | 8. Yamane I, Murakami O: 6,8-Dihydroxypurine: a novel growth factor for mammalian cells in vitro, isolated from a commercial peptone. J Cell Physiol. 1973 Apr;81(2):281-4. | | 9. Ohe T, Watanabe Y: Purification and properties of xanthine dehydrogenase from Streptomyces cyanogenus. J Biochem. 1979 Jul;86(1):45-53. | | 10. Simmonds HA, Sneddon W: Identification of 6,8-dihydroxypurine in the urine of a gouty patient treated with allopurinol. Clin Chim Acta. 1970 Nov;30(2):421-7. | | 11. Middelhoven WJ, Hoogkamer-Te Niet MC, Kreger-Van Rij NJ: Trichosporon adeninovorans sp. nov., a yeast species utilizing adenine, xanthine, uric acid, putrescine and primary n-alkylamines as the sole source of carbon, nitrogen and energy. Antonie Van Leeuwenhoek. 1984;50(4):369-78. | | 12. Durre P, Andreesen JR: Purine and glycine metabolism by purinolytic clostridia. J Bacteriol. 1983 Apr;154(1):192-9. | | 13. Kraushar MF, Nussbaum P, Kisch AL: Anaphylactic reaction to intravitreal cefazolin. Retina. 1994;14(2):187-8. | | 14. Keuzenkamp-Jansen CW, van Baal JM, De Abreu RA, de Jong JG, Zuiderent R, Trijbels JM: Detection and identification of 6-methylmercapto-8-hydoxypurine, a major metabolite of 6-mercaptopurine, in plasma during intravenous administration. Clin Chem. 1996 Mar;42(3):380-6. | | 15. Morris GS, Simmonds HA, Davies PM: Use of biological fluids for the rapid diagnosis of potentially lethal inherited disorders of human purine and pyrimidine metabolism. Biomed Chromatogr. 1986 Jun;1(3):109-18. | | 16. Yang TH, Hu ML: Intracellular levels of S-adenosylhomocysteine but not homocysteine are highly correlated to the expression of nm23-H1 and the level of 5-methyldeoxycytidine in human hepatoma cells with different invasion activities. Nutr Cancer. 2006;55(2):224-31. | | 17. Liu Z, Liu S, Xie Z, Blum W, Perrotti D, Paschka P, Klisovic R, Byrd J, Chan KK, Marcucci G: Characterization of in vitro and in vivo hypomethylating effects of decitabine in acute myeloid leukemia by a rapid, specific and sensitive LC-MS/MS method. Nucleic Acids Res. 2007;35(5):e31. Epub 2007 Jan 30. | | 18. Cho SH, Jung BH, Lee SH, Lee WY, Kong G, Chung BC: Direct determination of nucleosides in the urine of patients with breast cancer using column-switching liquid chromatography-tandem mass spectrometry. Biomed Chromatogr. 2006 Nov;20(11):1229-36. |
|
|---|