| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 03:19:56 UTC |
|---|
| Update Date | 2016-11-09 01:19:27 UTC |
|---|
| Accession Number | CHEM032357 |
|---|
| Identification |
|---|
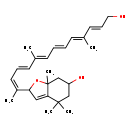
| Common Name | 5,8-Epoxy-5,8-dihydro-10'-apo-b,y-carotene-3,10'-diol |
|---|
| Class | Small Molecule |
|---|
| Description | 5,8-Epoxy-5,8-dihydro-10'-apo-b,y-carotene-3,10'-diol is found in citrus. 5,8-Epoxy-5,8-dihydro-10'-apo-b,y-carotene-3,10'-diol is isolated from Persea americana (avocado) and from Valencia orange juice. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | Not Available |
|---|
| Chemical Formula | C27H38O3 |
|---|
| Average Molecular Mass | 410.589 g/mol |
|---|
| Monoisotopic Mass | 410.282 g/mol |
|---|
| CAS Registry Number | 53905-15-0 |
|---|
| IUPAC Name | 2-[(2Z,4E,6E,8E,10E,12E)-14-hydroxy-6,11-dimethyltetradeca-2,4,6,8,10,12-hexaen-2-yl]-4,4,7a-trimethyl-2,4,5,6,7,7a-hexahydro-1-benzofuran-6-ol |
|---|
| Traditional Name | 2-[(2Z,4E,6E,8E,10E,12E)-14-hydroxy-6,11-dimethyltetradeca-2,4,6,8,10,12-hexaen-2-yl]-4,4,7a-trimethyl-2,5,6,7-tetrahydro-1-benzofuran-6-ol |
|---|
| SMILES | C\C(\C=C\CO)=C/C=C/C=C(\C)/C=C/C=C(/C)C1OC2(C)CC(O)CC(C)(C)C2=C1 |
|---|
| InChI Identifier | InChI=1S/C27H38O3/c1-20(11-7-8-12-21(2)14-10-16-28)13-9-15-22(3)24-17-25-26(4,5)18-23(29)19-27(25,6)30-24/h7-15,17,23-24,28-29H,16,18-19H2,1-6H3/b8-7+,13-9+,14-10+,20-11+,21-12+,22-15- |
|---|
| InChI Key | OOJOUQLXWXASLN-TVICGMDOSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as sesterterpenoids. These are terpenes composed of five consecutive isoprene units. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Sesterterpenoids |
|---|
| Direct Parent | Sesterterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Sesterterpenoid
- Long chain fatty alcohol
- Benzofuran
- Fatty alcohol
- Fatty acyl
- Cyclic alcohol
- Dihydrofuran
- Secondary alcohol
- Dialkyl ether
- Oxacycle
- Ether
- Organoheterocyclic compound
- Alcohol
- Organic oxygen compound
- Hydrocarbon derivative
- Primary alcohol
- Organooxygen compound
- Aliphatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aliphatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0udv-1409000000-eb429ad7cfc59d1f012b | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-000l-2141490000-7cd28af2be2d54de3f05 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-01ox-0129200000-ff8d7d4f182c31961d58 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0f9x-2945000000-35fad5ed96c103c1a70b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-05o0-7934000000-23a9c6723b3e64af3047 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0015900000-7c1c030a8a119325d2f9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4l-1219400000-070bdb65fa7ef1e302a1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-3649000000-5b13fd443c0189836066 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014i-0009100000-cb3fa5ef5e5f5596324b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-066r-0009300000-fb4591a8d574b299ce01 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-00ba-0039000000-a4c1d707b642f932ab80 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-02di-1209200000-a6152a27f3928816445a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4j-1937100000-5d32893ca7118f2aa190 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a5c-2900000000-e810e2854dfce1206755 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0039020 |
|---|
| FooDB ID | FDB018515 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 35014719 |
|---|
| ChEBI ID | 172652 |
|---|
| PubChem Compound ID | 131752516 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|