| Identification |
|---|
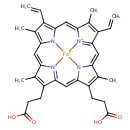
| Common Name | Haem |
|---|
| Class | Small Molecule |
|---|
| Description | |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| [3,7,12,17-Tetramethyl-8,13-divinylporphyrin-2,18-dipropanoato(2-)]iron(II) | ChEBI | | [Fe(ppix)] | ChEBI | | Fe(ppix) | ChEBI | | Ferroprotoheme | ChEBI | | Ferroprotoporphyrin IX | ChEBI | | Ferrous protoheme | ChEBI | | Ferrous protoheme IX | ChEBI | | Haem | ChEBI | | Iron(II) protoporphyrin IX | ChEBI | | Protoferroheme | ChEBI | | Protoheme | ChEBI | | Heme b | Kegg | | Protoheme IX | Kegg | | Heme | ChEBI |
|
|---|
| Chemical Formula | C34H32FeN4O4 |
|---|
| Average Molecular Mass | 616.487 g/mol |
|---|
| Monoisotopic Mass | 616.177 g/mol |
|---|
| CAS Registry Number | 14875-96-8 |
|---|
| IUPAC Name | 4,20-bis(2-carboxyethyl)-10,15-diethenyl-5,9,14,19-tetramethyl-2lambda5,22,23lambda5,25-tetraaza-1-ferraoctacyclo[11.9.1.1^{1,8}.1^{3,21}.0^{2,6}.0^{16,23}.0^{18,22}.0^{11,25}]pentacosa-2,4,6,8,10,12,14,16(23),17,19,21(24)-undecaene-2,23-bis(ylium)-1,1-diuide |
|---|
| Traditional Name | 4,20-bis(2-carboxyethyl)-10,15-diethenyl-5,9,14,19-tetramethyl-2lambda5,22,23lambda5,25-tetraaza-1-ferraoctacyclo[11.9.1.1^{1,8}.1^{3,21}.0^{2,6}.0^{16,23}.0^{18,22}.0^{11,25}]pentacosa-2,4,6,8,10,12,14,16(23),17,19,21(24)-undecaene-2,23-bis(ylium)-1,1-diuide |
|---|
| SMILES | CC1=C(CCC(O)=O)C2=CC3=[N]4C(=CC5=C(C)C(C=C)=C6C=C7C(C)=C(C=C)C8=[N]7[Fe]4(N2C1=C8)N56)C(C)=C3CCC(O)=O |
|---|
| InChI Identifier | InChI=1S/C34H34N4O4.Fe/c1-7-21-17(3)25-13-26-19(5)23(9-11-33(39)40)31(37-26)16-32-24(10-12-34(41)42)20(6)28(38-32)15-30-22(8-2)18(4)27(36-30)14-29(21)35-25;/h7-8,13-16H,1-2,9-12H2,3-6H3,(H4,35,36,37,38,39,40,41,42);/q;+2/p-2/b25-13-,26-13-,27-14-,28-15-,29-14-,30-15-,31-16-,32-16-; |
|---|
| InChI Key | KABFMIBPWCXCRK-RGGAHWMASA-L |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS ("Heme,1TMS,#1" TMS) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-066r-0000095000-501c4a776d38d61ab89a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0gi4-0000092000-64c21d0a0f8daa069fb9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03gs-4000590000-15136ce22e51bdc57377 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014i-0000049000-69ee302652cebc40f13b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-00r2-1000091000-d178bb83079a203a48dc | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-1000090000-f6e437751fbc08f12d1f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014j-0000059000-0ab427d87ba10992d498 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-014i-0000098000-650f7d2f023028ff2319 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-06fr-0000090000-2dff1da74eb12a246237 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-014j-0000079000-990502ea40621d9cad46 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-05ns-0000091000-3547d5c809ccf8987200 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0cdr-0000092000-e55d1ab39076367d5b9e | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum |
|
|---|