| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 01:58:37 UTC |
|---|
| Update Date | 2016-11-09 01:19:05 UTC |
|---|
| Accession Number | CHEM030549 |
|---|
| Identification |
|---|
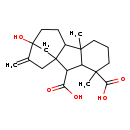
| Common Name | Gibberellin A53 |
|---|
| Class | Small Molecule |
|---|
| Description | Isolated from Vicia faba and spinach (Spinacia oleracea). Gibberellin A53 is found in many foods, some of which are sapodilla, cowpea, sorghum, and garden tomato. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| GA53 | HMDB | | Gibberellin 53 | HMDB |
|
|---|
| Chemical Formula | C20H28O5 |
|---|
| Average Molecular Mass | 348.433 g/mol |
|---|
| Monoisotopic Mass | 348.194 g/mol |
|---|
| CAS Registry Number | 51576-08-0 |
|---|
| IUPAC Name | 12-hydroxy-4,8-dimethyl-13-methylidenetetracyclo[10.2.1.0^{1,9}.0^{3,8}]pentadecane-2,4-dicarboxylic acid |
|---|
| Traditional Name | 12-hydroxy-4,8-dimethyl-13-methylidenetetracyclo[10.2.1.0^{1,9}.0^{3,8}]pentadecane-2,4-dicarboxylic acid |
|---|
| SMILES | [H][C@@]12CC[C@]3(O)C[C@]1(CC3=C)[C@@H](C(O)=O)[C@@]1([H])[C@@]2(C)CCC[C@@]1(C)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C20H28O5/c1-11-9-19-10-20(11,25)8-5-12(19)17(2)6-4-7-18(3,16(23)24)14(17)13(19)15(21)22/h12-14,25H,1,4-10H2,2-3H3,(H,21,22)(H,23,24)/t12-,13+,14-,17-,18+,19-,20-/m0/s1 |
|---|
| InChI Key | CZEMYYICWZPENF-VOLTXKGXSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as c20-gibberellin 6-carboxylic acids. These are c20-gibberellins with a carboxyl group at the 6-position. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Diterpenoids |
|---|
| Direct Parent | C20-gibberellin 6-carboxylic acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Gibberellane-6-carboxylic acid
- Dicarboxylic acid or derivatives
- Tertiary alcohol
- Cyclic alcohol
- Carboxylic acid
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Alcohol
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0fz9-0219000000-a3a32b5b1958eaf43854 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (3 TMS) - 70eV, Positive | splash10-002b-4021790000-be27b9ca49c8b719a502 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-0029000000-c96aeeb9e91b4028c07a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-01q0-0179000000-9a1ffa403e54886ddd84 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-052r-2692000000-2028bd409181d9869708 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0f6t-0029000000-4b508b38191430bae1f1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0zg1-0079000000-3cb7e1465f14fd5a603f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4r-1094000000-ba4002cfdf21089dc56e | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0036895 |
|---|
| FooDB ID | FDB015855 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00000053 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 4420151 |
|---|
| ChEBI ID | 27433 |
|---|
| PubChem Compound ID | 5253706 |
|---|
| Kegg Compound ID | C06094 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|