| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 01:45:16 UTC |
|---|
| Update Date | 2016-11-09 01:19:02 UTC |
|---|
| Accession Number | CHEM030234 |
|---|
| Identification |
|---|
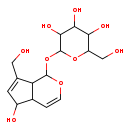
| Common Name | Aucubin |
|---|
| Class | Small Molecule |
|---|
| Description | Aucubin is found in common verbena. Aucubin is a monoterpenoid based compound. Aucubin, like all iridoids, has a cyclopentan-[C]-pyran skeleton. Iridoids can consist of ten, nine, or rarely eight carbons in which C11 is more frequently missing than C10. Aucubin has 10 carbons with the C11 carbon missing. The stereochemical configurations at C5 and C9 lead to cis fused rings, which are common to all iridoids containing carbocylclic- or seco-skeleton in non-rearranged form. Oxidative cleavage at C7-C8 bond affords secoiridoids. The last steps in the biosynthesis of iridoids usually consist of O-glycosylation and O-alkylation. Aucubin, a glycoside iridoid, has an O-linked glucose moiety. Aucubin is an iridoid glycoside. Iridoids are commonly found in plants and function as defensive compounds. Irioids decrease the growth rates of many generalist herbivores. Aucubin is found in the leaves of Aucuba japonica (Cornaceae), Eucommia ulmoides (Eucommiaceae), and Plantago asiatic (Plantaginaceae), etc, plants used in traditional Chinese and folk medicine. Aucubin was found to protect against liver damage induced by carbon tetrachloride or alpha-amanitin in mice and rats when 80 mg/kg was dosed intraperitoneally. Geranyl pyrophosphate is the precursor for iridoids. Geranyl phosphate is generated through the mevalonate pathway or the methylerythritol phosphate pathway. The initial steps of the pathway involve the fusion of three molecules of acetyl-CoA to produce the C6 compound 3-hydroxy-3-methylglutaryl-CoA (HMG-CoA). HMG-CoA is then reduced in two steps by the enzyme HMG-CoA reductase. The resulting mevalonate is then sequentially phosphorylated by two separate kinases, mevalonate kinase and phosphomevalonate kinase, to form 5-pyrophosphomevalonate. Phosphosphomevalonate decarboxylase through a concerted decarboxylation reaction affords isopentenyl pyrophosphate (IPP). IPP is the basic C5 building block that is added to prenyl phosphate cosubstrates to form longer chains. IPP is isomerized to the allylic ester dimethylallyl pyrophosphate (DMAPP) by IPP isomerase. Through a multistep process, including the dephosphorylation DMAPP, IPP and DMAPP are combinded to from the C10 compound geranyl pyrophosphate (GPP). Geranyl pyrophosphate is a major branch point for terpenoid synthesis. The cyclizaton reaction to form the iridoid pyrane ring may result from one of two routes: route 1 - a hydride nucleophillic attack on C1 will lead to 1-O-carbonyl atom attack on C3, yielding the lactone ring; route 2 - loss of proton from carbon 4 leads to the formation of a double bond C3-C4; consequently the 3-0-carbonyl atom will attach to C1 |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| Aucuboside | HMDB | | Rhimantin | HMDB | | Rhinanthin | HMDB |
|
|---|
| Chemical Formula | C15H22O9 |
|---|
| Average Molecular Mass | 346.330 g/mol |
|---|
| Monoisotopic Mass | 346.126 g/mol |
|---|
| CAS Registry Number | 479-98-1 |
|---|
| IUPAC Name | 2-{[5-hydroxy-7-(hydroxymethyl)-1H,4aH,5H,7aH-cyclopenta[c]pyran-1-yl]oxy}-6-(hydroxymethyl)oxane-3,4,5-triol |
|---|
| Traditional Name | aucubin |
|---|
| SMILES | OCC1OC(OC2OC=CC3C(O)C=C(CO)C23)C(O)C(O)C1O |
|---|
| InChI Identifier | InChI=1S/C15H22O9/c16-4-6-3-8(18)7-1-2-22-14(10(6)7)24-15-13(21)12(20)11(19)9(5-17)23-15/h1-3,7-21H,4-5H2 |
|---|
| InChI Key | RJWJHRPNHPHBRN-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as iridoid o-glycosides. These are iridoid monoterpenes containing a glycosyl (usually a pyranosyl) moiety linked to the iridoid skeleton. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Terpene glycosides |
|---|
| Direct Parent | Iridoid O-glycosides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Iridoid o-glycoside
- Hexose monosaccharide
- Glycosyl compound
- Iridoid-skeleton
- O-glycosyl compound
- Bicyclic monoterpenoid
- Monoterpenoid
- Monosaccharide
- Oxane
- Secondary alcohol
- Acetal
- Oxacycle
- Organoheterocyclic compound
- Polyol
- Alcohol
- Hydrocarbon derivative
- Primary alcohol
- Organic oxygen compound
- Organooxygen compound
- Aliphatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aliphatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00or-4719000000-a03c5ea19b82d7991f35 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (4 TMS) - 70eV, Positive | splash10-01b9-2252049000-fb4fdbe14aa2fef93db3 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-02vr-0904000000-0dc8cc1fee5ec1d7b66a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-014i-0901000000-23ae99445d0f902ebb10 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-014i-1900000000-0197bf91c00175908c8f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-00nb-0917000000-5b5a2c303dee5ed5b778 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0159-1901000000-dcbc777e43693cc64c6f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-001l-5900000000-66eed55623a37e730cdc | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0002-0209000000-6d1f85d0258419cba6e0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-005a-5925000000-7b6e750efe6aab6eda8d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0uxr-6910000000-9a98e60322aa35e884b0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00mk-0709000000-9b62837c71245c712dd9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00kb-0901000000-8bd04921ef01bfdb1bb4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-07bs-4900000000-1f7344e04d4a474d0b71 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0036562 |
|---|
| FooDB ID | FDB015466 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00003073 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Aucubin |
|---|
| Chemspider ID | 308989 |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 348157 |
|---|
| Kegg Compound ID | C09771 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Hung JY, Yang CJ, Tsai YM, Huang HW, Huang MS: Antiproliferative activity of aucubin is through cell cycle arrest and apoptosis in human non-small cell lung cancer A549 cells. Clin Exp Pharmacol Physiol. 2008 Sep;35(9):995-1001. doi: 10.1111/j.1440-1681.2008.04935.x. Epub 2008 Apr 21. | | 2. Simons K, Toomre D: Lipid rafts and signal transduction. Nat Rev Mol Cell Biol. 2000 Oct;1(1):31-9. | | 3. Watson AD: Thematic review series: systems biology approaches to metabolic and cardiovascular disorders. Lipidomics: a global approach to lipid analysis in biological systems. J Lipid Res. 2006 Oct;47(10):2101-11. Epub 2006 Aug 10. | | 4. Sethi JK, Vidal-Puig AJ: Thematic review series: adipocyte biology. Adipose tissue function and plasticity orchestrate nutritional adaptation. J Lipid Res. 2007 Jun;48(6):1253-62. Epub 2007 Mar 20. | | 5. Lingwood D, Simons K: Lipid rafts as a membrane-organizing principle. Science. 2010 Jan 1;327(5961):46-50. doi: 10.1126/science.1174621. | | 6. Yannai, Shmuel. (2004) Dictionary of food compounds with CD-ROM: Additives, flavors, and ingredients. Boca Raton: Chapman & Hall/CRC. | | 7. The lipid handbook with CD-ROM |
|
|---|