| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 01:33:07 UTC |
|---|
| Update Date | 2016-11-09 01:18:59 UTC |
|---|
| Accession Number | CHEM029964 |
|---|
| Identification |
|---|
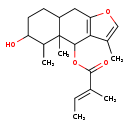
| Common Name | 6-Angeloylfuranofukinol |
|---|
| Class | Small Molecule |
|---|
| Description | 6-Angeloylfuranofukinol is found in green vegetables. 6-Angeloylfuranofukinol is a constituent of Petasites japonicus (sweet coltsfoot) |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 6-Hydroxy-3,4a,5-trimethyl-4H,4ah,5H,6H,7H,8H,8ah,9H-naphtho[2,3-b]furan-4-yl (2E)-2-methylbut-2-enoic acid | HMDB |
|
|---|
| Chemical Formula | C20H28O4 |
|---|
| Average Molecular Mass | 332.434 g/mol |
|---|
| Monoisotopic Mass | 332.199 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | 6-hydroxy-3,4a,5-trimethyl-4H,4aH,5H,6H,7H,8H,8aH,9H-naphtho[2,3-b]furan-4-yl (2E)-2-methylbut-2-enoate |
|---|
| Traditional Name | 6-hydroxy-3,4a,5-trimethyl-4H,5H,6H,7H,8H,8aH,9H-naphtho[2,3-b]furan-4-yl (2E)-2-methylbut-2-enoate |
|---|
| SMILES | C\C=C(/C)C(=O)OC1C2=C(CC3CCC(O)C(C)C13C)OC=C2C |
|---|
| InChI Identifier | InChI=1S/C20H28O4/c1-6-11(2)19(22)24-18-17-12(3)10-23-16(17)9-14-7-8-15(21)13(4)20(14,18)5/h6,10,13-15,18,21H,7-9H2,1-5H3/b11-6+ |
|---|
| InChI Key | QSXNOUPYXMWUKT-IZZDOVSWSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as eremophilane, 8,9-secoeremophilane and furoeremophilane sesquiterpenoids. These are sesquiterpenoids with a structure based either on the eremophilane skeleton, its 8,9-seco derivative, or the furoeremophilane skeleton. Eremophilanes have been shown to be derived from eudesmanes by migration of the methyl group at C-10 to C-5. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Sesquiterpenoids |
|---|
| Direct Parent | Eremophilane, 8,9-secoeremophilane and furoeremophilane sesquiterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Furoeremophilane sesquiterpenoid
- Naphthofuran
- Benzofuran
- Fatty acid ester
- Fatty acyl
- Cyclic alcohol
- Heteroaromatic compound
- Furan
- Alpha,beta-unsaturated carboxylic ester
- Enoate ester
- Carboxylic acid ester
- Secondary alcohol
- Carboxylic acid derivative
- Oxacycle
- Organoheterocyclic compound
- Monocarboxylic acid or derivatives
- Hydrocarbon derivative
- Organic oxide
- Alcohol
- Organic oxygen compound
- Carbonyl group
- Organooxygen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-05o0-9053000000-be01ea31423dfe805803 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-0569-9225000000-910ac19c39e5979dbf9a | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00lr-2039000000-2b969d352b2c0472963b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-001i-9562000000-5eebc899456b45e0d658 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0kai-9210000000-0e69aad016e0a7b82b55 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-1019000000-609104691c0fca84352a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-7059000000-4e04a0087e62ad93ddd6 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-05nb-7290000000-86b5266725199b58e795 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001j-7039000000-f71ac89f2443e259a1a0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001j-9056000000-9a79012566eb09266356 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0pb9-9000000000-aa5880a0b8fe9ddffe66 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00lr-0094000000-626cad1dbd0e012997c2 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0159-0194000000-2b6fa55833db6793f0aa | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-8491000000-ae31a375e4155bf3f6e6 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0036146 |
|---|
| FooDB ID | FDB014997 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 4900481 |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 6374435 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|