| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 01:22:06 UTC |
|---|
| Update Date | 2016-11-09 01:18:56 UTC |
|---|
| Accession Number | CHEM029724 |
|---|
| Identification |
|---|
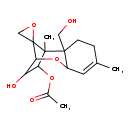
| Common Name | 4-Acetoxyscirpene-3,15-diol |
|---|
| Class | Small Molecule |
|---|
| Description | 4-Acetoxyscirpene-3,15-diol is a mycotoxin from Fusarium roseum and Fusarium sulphureu |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 4beta-Acetoxyscirpene-3alpha,15-diol | MeSH | | 4-Acetoxyscirpenediol | MeSH | | 10'-Hydroxy-2'-(hydroxymethyl)-1',5'-dimethyl-8'-oxaspiro[oxirane-2,12'-tricyclo[7.2.1.0²,⁷]dodecan]-5'-en-11'-yl acetic acid | Generator | | 4-Acetoxyscirpene-3,15-diol | MeSH |
|
|---|
| Chemical Formula | C17H24O6 |
|---|
| Average Molecular Mass | 324.369 g/mol |
|---|
| Monoisotopic Mass | 324.157 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | 10'-hydroxy-2'-(hydroxymethyl)-1',5'-dimethyl-8'-oxaspiro[oxirane-2,12'-tricyclo[7.2.1.0²,⁷]dodecan]-5'-en-11'-yl acetate |
|---|
| Traditional Name | 10'-hydroxy-2'-(hydroxymethyl)-1',5'-dimethyl-8'-oxaspiro[oxirane-2,12'-tricyclo[7.2.1.0²,⁷]dodecan]-5'-en-11'-yl acetate |
|---|
| SMILES | CC(=O)OC1C(O)C2OC3C=C(C)CCC3(CO)C1(C)C21CO1 |
|---|
| InChI Identifier | InChI=1S/C17H24O6/c1-9-4-5-16(7-18)11(6-9)23-14-12(20)13(22-10(2)19)15(16,3)17(14)8-21-17/h6,11-14,18,20H,4-5,7-8H2,1-3H3 |
|---|
| InChI Key | YIFMFXZUYBFVFL-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as trichothecenes. These are sesquiterpene mycotoxins structurally characterized by the presence of an epoxide ring and a benzopyran derivative with a variant number of hydroxyl, acetyl, or other substituents. The most important structural features causing the biological activities of trichothecenes are the 12,13-epoxy ring, the presence of hydroxyl or acetyl groups at appropriate positions on the trichothecene nucleus and the structure and position of the side-chain. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Sesquiterpenoids |
|---|
| Direct Parent | Trichothecenes |
|---|
| Alternative Parents | |
|---|
| Substituents | - Trichothecene skeleton
- Oxepane
- Oxane
- Cyclic alcohol
- Carboxylic acid ester
- Secondary alcohol
- Oxacycle
- Carboxylic acid derivative
- Organoheterocyclic compound
- Dialkyl ether
- Oxirane
- Ether
- Monocarboxylic acid or derivatives
- Carbonyl group
- Alcohol
- Organooxygen compound
- Organic oxygen compound
- Organic oxide
- Primary alcohol
- Hydrocarbon derivative
- Aliphatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aliphatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00di-1971000000-672a74e68c69b4aa0864 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-0007-4391100000-ddd845e5c8c9e765142e | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-05r0-0069000000-edbf95134c169781fc5b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0aor-1494000000-0f562d03c0ed6b44b675 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-014i-9470000000-8447aa4f0c47a4cff2e7 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-05fr-3069000000-e075d5b14c12b589d499 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0bu3-3393000000-d3460f4398479d1ba190 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-5900000000-485823559d284aa43163 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-05fr-8009000000-665c3440a41beb01d455 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-9000000000-b183e5b2445ad7ad0405 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-052f-5092000000-1b99ee87c880f62ac330 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-004i-0009000000-f933cd2781eb4911e709 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-004j-0097000000-6a3f346078033195ab23 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0pb9-9540000000-14d6c1da5207fcc49238 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0035847 |
|---|
| FooDB ID | FDB014625 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 3242824 |
|---|
| ChEBI ID | 175143 |
|---|
| PubChem Compound ID | 4025704 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|