| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 01:01:23 UTC |
|---|
| Update Date | 2016-11-09 01:18:50 UTC |
|---|
| Accession Number | CHEM029224 |
|---|
| Identification |
|---|
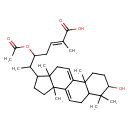
| Common Name | Ganoderic acid S |
|---|
| Class | Small Molecule |
|---|
| Description | Ganoderic acid S is found in mushrooms. Ganoderic acid S is a constituent of cultured mycelium of Ganoderma lucidum (reishi). |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| Ganoderate S | Generator | | 6a-Carissanol | HMDB | | 6Α-carissanol | HMDB | | (+)-6alpha-Carissanol | HMDB | | [4AS-(4aalpha,7alpha,8beta)]-4,4a,5,6,7,8-hexahydro-8-hydroxy-7-(1-hydroxy-1-methylethyl)-1,4a-dimethyl-2(3H)-naphthalenone | HMDB | | (2E)-5-(Acetyloxy)-6-{5-hydroxy-2,6,6,11,15-pentamethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadeca-1(17),9-dien-14-yl}-2-methylhept-2-enoate | HMDB |
|
|---|
| Chemical Formula | C32H48O5 |
|---|
| Average Molecular Mass | 512.721 g/mol |
|---|
| Monoisotopic Mass | 512.350 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | (2E)-5-(acetyloxy)-6-{5-hydroxy-2,6,6,11,15-pentamethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadeca-1(17),9-dien-14-yl}-2-methylhept-2-enoic acid |
|---|
| Traditional Name | (2E)-5-(acetyloxy)-6-{5-hydroxy-2,6,6,11,15-pentamethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadeca-1(17),9-dien-14-yl}-2-methylhept-2-enoic acid |
|---|
| SMILES | CC(C(C\C=C(/C)C(O)=O)OC(C)=O)C1CCC2(C)C3=CCC4C(C)(C)C(O)CCC4(C)C3=CCC12C |
|---|
| InChI Identifier | InChI=1S/C32H48O5/c1-19(28(35)36)9-11-25(37-21(3)33)20(2)22-13-17-32(8)24-10-12-26-29(4,5)27(34)15-16-30(26,6)23(24)14-18-31(22,32)7/h9-10,14,20,22,25-27,34H,11-13,15-18H2,1-8H3,(H,35,36)/b19-9+ |
|---|
| InChI Key | FRRMMEJOPQSLSE-DJKKODMXSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as triterpenoids. These are terpene molecules containing six isoprene units. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Triterpenoids |
|---|
| Direct Parent | Triterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Triterpenoid
- Cholane-skeleton
- Hydroxy bile acid, alcohol, or derivatives
- Monohydroxy bile acid, alcohol, or derivatives
- Bile acid, alcohol, or derivatives
- Steroid ester
- Steroid acid
- 3-hydroxy-delta-7-steroid
- 3-hydroxysteroid
- Hydroxysteroid
- Delta-7-steroid
- Steroid
- Medium-chain fatty acid
- Hydroxy fatty acid
- Unsaturated fatty acid
- Fatty acid
- Dicarboxylic acid or derivatives
- Fatty acyl
- Cyclic alcohol
- Secondary alcohol
- Carboxylic acid ester
- Carboxylic acid derivative
- Carboxylic acid
- Organooxygen compound
- Carbonyl group
- Hydrocarbon derivative
- Alcohol
- Organic oxygen compound
- Organic oxide
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-01pn-1001900000-54288b5e89578dd7a15e | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-0006-2103139000-e5a9e46017bd7ae66284 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-01ot-0000910000-cbb25a22a50983dde5a4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0f72-0001900000-b460379fb861e9648885 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-11ba-0013900000-f275c1b2801a2adc639f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-03di-2000970000-871d5813a819254e8f64 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0n2c-3000910000-d8b6de2682280c379219 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-052f-7004900000-bf591c34e488a0c2fe51 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0903810000-1f65eb5e9d6809ba1ee8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a7i-1414900000-22c0864466ff3744a935 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0903-6798000000-1632832ad9868f9d92be | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-6000930000-028d849c4650dd2feeef | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-9000200000-0e94b1c127887e27d559 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4l-9201100000-ec0e872594f90c8a7715 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0035313 |
|---|
| FooDB ID | FDB013982 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00012701 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | Not Available |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 13916698 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|