| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 00:54:03 UTC |
|---|
| Update Date | 2016-11-09 01:18:48 UTC |
|---|
| Accession Number | CHEM029066 |
|---|
| Identification |
|---|
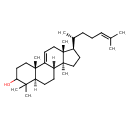
| Common Name | Parkeol |
|---|
| Class | Small Molecule |
|---|
| Description | Parkeol is found in fats and oils. Parkeol is a constituent of Butyrospermum parkii (shea) |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 4,4,14alpha-Trimethyl-5alpha-cholesta-9(11),24-dien-3beta-ol | HMDB | | 5alpha-Lanosta-9(11),24-dien-3beta-ol | HMDB | | Delta(9(11),24)-Lanostadien-3beta-ol | HMDB | | Lanost-9(11)-en-3beta-ol | HMDB | | 4,4,14a-Trimethyl-5a-cholesta-9(11),24-dien-3b-ol | HMDB | | 4,4,14Α-trimethyl-5α-cholesta-9(11),24-dien-3β-ol | HMDB | | 5a-Lanosta-9(11),24-dien-3b-ol | HMDB | | 5Α-lanosta-9(11),24-dien-3β-ol | HMDB | | delta(9(11),24)-Lanostadien-3b-ol | HMDB | | Δ(9(11),24)-lanostadien-3β-ol | HMDB | | Lanost-9(11)-en-3b-ol | HMDB | | Lanost-9(11)-en-3β-ol | HMDB | | Lanost-9(11)-en-3 beta-ol | HMDB | | Lanost-9(11)-en-3-ol | HMDB |
|
|---|
| Chemical Formula | C30H50O |
|---|
| Average Molecular Mass | 426.717 g/mol |
|---|
| Monoisotopic Mass | 426.386 g/mol |
|---|
| CAS Registry Number | 514-45-4 |
|---|
| IUPAC Name | (2S,7R,10S,11S,14R,15R)-2,6,6,11,15-pentamethyl-14-[(2R)-6-methylhept-5-en-2-yl]tetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadec-1(17)-en-5-ol |
|---|
| Traditional Name | (2S,7R,10S,11S,14R,15R)-2,6,6,11,15-pentamethyl-14-[(2R)-6-methylhept-5-en-2-yl]tetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadec-1(17)-en-5-ol |
|---|
| SMILES | [H][C@@]12CC[C@@]3([H])C(C)(C)C(O)CC[C@]3(C)C1=CC[C@]1(C)[C@H](CC[C@@]21C)[C@H](C)CCC=C(C)C |
|---|
| InChI Identifier | InChI=1S/C30H50O/c1-20(2)10-9-11-21(3)22-14-18-30(8)24-12-13-25-27(4,5)26(31)16-17-28(25,6)23(24)15-19-29(22,30)7/h10,15,21-22,24-26,31H,9,11-14,16-19H2,1-8H3/t21-,22-,24-,25+,26?,28-,29-,30+/m1/s1 |
|---|
| InChI Key | MLVSYGCURCOSKP-LFXYTPDNSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as triterpenoids. These are terpene molecules containing six isoprene units. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Triterpenoids |
|---|
| Direct Parent | Triterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Triterpenoid
- Cholesterol-skeleton
- Cholestane-skeleton
- 14-alpha-methylsteroid
- Hydroxysteroid
- 3-hydroxysteroid
- Steroid
- Cyclic alcohol
- Secondary alcohol
- Organic oxygen compound
- Hydrocarbon derivative
- Organooxygen compound
- Alcohol
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-03dj-1019400000-4684ce900e2bdc746072 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-00lr-3001900000-8451a437af3bdb2462bf | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a6r-0002900000-2cfcf7ce4b5b21140eae | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0pdi-3149500000-61a41ba954dca6136093 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0uxr-4249100000-2bb374a801f5e66d3c9e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-0000900000-120a00ee008954da444d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-0000900000-017947c10e39f58df695 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4l-1009400000-afc7034885289079e0af | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-0000900000-590c9e4adfdd12b64d93 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-0000900000-e46a7ceb14ec0697db5a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-1001900000-67a4126d96742d377bfc | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a6u-9101200000-a16a2818c31140982176 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0aou-9102000000-1dfa95bffb6fb59ced74 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0aou-9501000000-2cb83003c3cd64db470c | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0035128 |
|---|
| FooDB ID | FDB013763 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00023785 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | CPD-13879 |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Parkeol |
|---|
| Chemspider ID | 23255009 |
|---|
| ChEBI ID | 63460 |
|---|
| PubChem Compound ID | 131751669 |
|---|
| Kegg Compound ID | C19918 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|