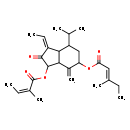| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 00:16:15 UTC |
|---|
| Update Date | 2016-11-09 01:18:37 UTC |
|---|
| Accession Number | CHEM028176 |
|---|
| Identification |
|---|
| Common Name | 2-Angeloyl-9-(3-methyl-2E-pentenoyl)-2b,9a-dihydroxy-4Z,10(14)-oplopadien-3-one |
|---|
| Class | Small Molecule |
|---|
| Description | 2-Angeloyl-9-(3-methyl-2E-pentenoyl)-2b,9a-dihydroxy-4Z,10(14)-oplopadien-3-one is found in tea. 2-Angeloyl-9-(3-methyl-2E-pentenoyl)-2b,9a-dihydroxy-4Z,10(14)-oplopadien-3-one is a constituent of Tussilago farfara (coltsfoot). |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| (1Z)-1-Ethylidene-3-{[(2Z)-2-methylbut-2-enoyl]oxy}-4-methylidene-2-oxo-7-(propan-2-yl)-octahydro-1H-inden-5-yl (2E)-3-methylpent-2-enoic acid | HMDB |
|
|---|
| Chemical Formula | C26H36O5 |
|---|
| Average Molecular Mass | 428.561 g/mol |
|---|
| Monoisotopic Mass | 428.256 g/mol |
|---|
| CAS Registry Number | 237407-02-2 |
|---|
| IUPAC Name | (1Z)-1-ethylidene-3-{[(2Z)-2-methylbut-2-enoyl]oxy}-4-methylidene-2-oxo-7-(propan-2-yl)-octahydro-1H-inden-5-yl (2E)-3-methylpent-2-enoate |
|---|
| Traditional Name | (1Z)-1-ethylidene-7-isopropyl-3-{[(2Z)-2-methylbut-2-enoyl]oxy}-4-methylidene-2-oxo-hexahydroinden-5-yl (2E)-3-methylpent-2-enoate |
|---|
| SMILES | CC\C(C)=C\C(=O)OC1CC(C(C)C)C2C(C(OC(=O)C(\C)=C/C)C(=O)\C2=C/C)C1=C |
|---|
| InChI Identifier | InChI=1S/C26H36O5/c1-9-15(6)12-21(27)30-20-13-19(14(4)5)23-18(11-3)24(28)25(22(23)17(20)8)31-26(29)16(7)10-2/h10-12,14,19-20,22-23,25H,8-9,13H2,1-7H3/b15-12+,16-10-,18-11- |
|---|
| InChI Key | FBGUGCFBJJKKCK-AHFXLWIPSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as sesquiterpenoids. These are terpenes with three consecutive isoprene units. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Sesquiterpenoids |
|---|
| Direct Parent | Sesquiterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Sesquiterpenoid
- Oplopane sesquiterpenoid
- Fatty acid ester
- Alpha-acyloxy ketone
- Dicarboxylic acid or derivatives
- Fatty acyl
- Enoate ester
- Alpha,beta-unsaturated carboxylic ester
- Cyclic ketone
- Ketone
- Carboxylic acid ester
- Carboxylic acid derivative
- Hydrocarbon derivative
- Organic oxide
- Organooxygen compound
- Organic oxygen compound
- Carbonyl group
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00lr-9014100000-6b9f26f0cd4cffb886e7 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-057i-9005400000-2072db475f0eb5e35f92 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0pb9-9001000000-8afc3c68b79f66fca1c1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0pc0-9010000000-2dc8a082782a36360329 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004j-6006900000-2aefce91c4b2036c867a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001j-8129300000-b1194f50974af2a460a9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0002-9041000000-72684d6070c19cf6dddc | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-014i-0091000000-16808289c7f74d3f33d0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0159-0090000000-a6cfd3a4ac9346f0c373 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-053u-7390000000-89ccb3236361288301f9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0002-9022300000-a0499d1d9f785aeed5d8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001j-5294100000-7a71f13ec14c61dede84 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a59-8191000000-7ee0d9622396be55f532 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0034064 |
|---|
| FooDB ID | FDB012313 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 35013693 |
|---|
| ChEBI ID | 175477 |
|---|
| PubChem Compound ID | 131751519 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|