| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-26 00:00:23 UTC |
|---|
| Update Date | 2016-11-09 01:18:32 UTC |
|---|
| Accession Number | CHEM027823 |
|---|
| Identification |
|---|
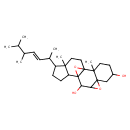
| Common Name | 5,6:8,9-Diepoxyergost-22-ene-3,7alpha-diol |
|---|
| Class | Small Molecule |
|---|
| Description | Constituent of a Rhei species Torachrysone 8-(6-oxalylglucoside) is found in green vegetables. |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 5,6:8,9-Diepoxy-24-methylcholest-22-ene-3,7-diol | HMDB | | 5,6:8,9-Diepoxyergost-22-ene-3,7-diol | HMDB | | 5,6:8,9-Diepoxyergost-22-ene-3,7b-diol | Generator | | 5,6:8,9-Diepoxyergost-22-ene-3,7β-diol | Generator |
|
|---|
| Chemical Formula | C28H44O4 |
|---|
| Average Molecular Mass | 444.647 g/mol |
|---|
| Monoisotopic Mass | 444.324 g/mol |
|---|
| CAS Registry Number | 243449-51-6 |
|---|
| IUPAC Name | 15-[(3E)-5,6-dimethylhept-3-en-2-yl]-2,16-dimethyl-8,19-dioxahexacyclo[9.7.1.0¹,¹¹.0²,⁷.0⁷,⁹.0¹²,¹⁶]nonadecane-5,10-diol |
|---|
| Traditional Name | 15-[(3E)-5,6-dimethylhept-3-en-2-yl]-2,16-dimethyl-8,19-dioxahexacyclo[9.7.1.0¹,¹¹.0²,⁷.0⁷,⁹.0¹²,¹⁶]nonadecane-5,10-diol |
|---|
| SMILES | CC(C)C(C)\C=C\C(C)C1CCC2C34OC3(CCC12C)C1(C)CCC(O)CC11OC1C4O |
|---|
| InChI Identifier | InChI=1S/C28H44O4/c1-16(2)17(3)7-8-18(4)20-9-10-21-24(20,5)13-14-27-25(6)12-11-19(29)15-26(25)23(31-26)22(30)28(21,27)32-27/h7-8,16-23,29-30H,9-15H2,1-6H3/b8-7+ |
|---|
| InChI Key | KAQBBVJKPGNKRN-BQYQJAHWSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as phenolic glycosides. These are organic compounds containing a phenolic structure attached to a glycosyl moiety. Some examples of phenolic structures include lignans, and flavonoids. Among the sugar units found in natural glycosides are D-glucose, L-Fructose, and L rhamnose. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Phenolic glycosides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Phenolic glycoside
- 1-naphthol
- O-glycosyl compound
- Naphthalene
- Acetophenone
- Anisole
- Aryl ketone
- Aryl alkyl ketone
- 1-hydroxy-4-unsubstituted benzenoid
- Alkyl aryl ether
- Benzenoid
- Oxane
- Dicarboxylic acid or derivatives
- Monosaccharide
- Vinylogous acid
- Secondary alcohol
- Carboxylic acid ester
- Ketone
- Organoheterocyclic compound
- Polyol
- Oxacycle
- Acetal
- Carboxylic acid derivative
- Ether
- Carboxylic acid
- Alcohol
- Hydrocarbon derivative
- Organic oxide
- Carbonyl group
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00or-4593700000-b3b569567f29e65a3eb8 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-00di-5360190000-0d354d65aaeea06db90a | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-004j-1002900000-892f5d9d943ef38308e8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0044-9008500000-23a63954d3d55d67b322 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-001r-9112000000-753f1a6ce34360f9307f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0006-0000900000-1dbd2cb93dbf86937cff | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-002f-0000900000-bd99ba62e765b33bf0f0 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-00b9-6169400000-e870124a9ddb35843fd3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0002-1003900000-917fe03485ef2bcfbddf | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a7j-9204600000-7789c3b20f94ebb4c611 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a59-9101000000-86bcc39f148713af817f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0006-0000900000-4a1a4016cf02e8b45f39 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0006-0000900000-4a1a4016cf02e8b45f39 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-0002900000-c5dcc770479bad9a7ac4 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0039135 |
|---|
| FooDB ID | FDB018653 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | Not Available |
|---|
| ChEBI ID | Not Available |
|---|
| PubChem Compound ID | 78385603 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|