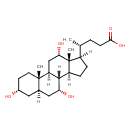
| Synonyms | | Value | Source |
|---|
| 3alpha,7alpha,12alpha-Trihydroxy-5alpha-cholanoic acid | ChEBI | | 5alpha-Cholic acid | ChEBI | | Allocholate | Kegg | | (3a,5a,7a,12a)-3,7,12-Trihydroxycholan-24-Oate | Generator | | (3a,5a,7a,12a)-3,7,12-Trihydroxycholan-24-Oic acid | Generator | | (3alpha,5alpha,7alpha,12alpha)-3,7,12-Trihydroxycholan-24-Oate | Generator | | (3Α,5α,7α,12α)-3,7,12-trihydroxycholan-24-Oate | Generator | | (3Α,5α,7α,12α)-3,7,12-trihydroxycholan-24-Oic acid | Generator | | 3a,7a,12a-Trihydroxy-5a-cholanoate | Generator | | 3a,7a,12a-Trihydroxy-5a-cholanoic acid | Generator | | 3alpha,7alpha,12alpha-Trihydroxy-5alpha-cholanoate | Generator | | 5a-Cholate | Generator | | 5a-Cholic acid | Generator | | 5alpha-Cholate | Generator | | (3alpha,5beta,7alpha,12alpha)-3,7,12-Trihydroxycholan-24-Oic acid | HMDB | | (3a,5b,7a,12a)-3,7,12-Trihydroxycholan-24-Oate | HMDB | | (3a,5b,7a,12a)-3,7,12-Trihydroxycholan-24-Oic acid | HMDB | | (3alpha,5beta,7alpha,12alpha)-3,7,12-Trihydroxycholan-24-Oate | HMDB | | (3Α,5β,7α,12α)-3,7,12-trihydroxycholan-24-Oate | HMDB | | (3Α,5β,7α,12α)-3,7,12-trihydroxycholan-24-Oic acid | HMDB | | 3a,7a,12a-Trihydroxy-5b-cholan-24-Oate | HMDB | | 3a,7a,12a-Trihydroxy-5b-cholan-24-Oic acid | HMDB | | 3a,7a,12a-Trihydroxycholanate | HMDB | | 3a,7a,12a-Trihydroxycholanic acid | HMDB | | 5alpha-Allocholic acid | HMDB | | 5Α-allocholic acid | HMDB | | Allocholic acid | HMDB |
|
|---|
| IUPAC Name | (4R)-4-[(1S,2S,5R,7R,9R,10R,11S,14R,15R,16S)-5,9,16-trihydroxy-2,15-dimethyltetracyclo[8.7.0.0^{2,7}.0^{11,15}]heptadecan-14-yl]pentanoic acid |
|---|
| Traditional Name | (4R)-4-[(1S,2S,5R,7R,9R,10R,11S,14R,15R,16S)-5,9,16-trihydroxy-2,15-dimethyltetracyclo[8.7.0.0^{2,7}.0^{11,15}]heptadecan-14-yl]pentanoic acid |
|---|