| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 23:53:04 UTC |
|---|
| Update Date | 2016-11-09 01:18:30 UTC |
|---|
| Accession Number | CHEM027649 |
|---|
| Identification |
|---|
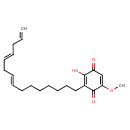
| Common Name | Sorgoleone 358 |
|---|
| Class | Small Molecule |
|---|
| Description | Sorgoleone 358 is found in cereals and cereal products. Sorgoleone 358 is a constituent of the etiolated seedlings of Sorghum bicolor (sorghum). |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 2-Hydroxy-5-methoxy-3-(8,11,14-pentadecatrienyl)-1,4-benzoquinone | HMDB | | Sorgoleone | HMDB |
|
|---|
| Chemical Formula | C22H30O4 |
|---|
| Average Molecular Mass | 358.471 g/mol |
|---|
| Monoisotopic Mass | 358.214 g/mol |
|---|
| CAS Registry Number | 105018-76-6 |
|---|
| IUPAC Name | 2-hydroxy-5-methoxy-3-[(8E,11E)-pentadeca-8,11,14-trien-1-yl]cyclohexa-2,5-diene-1,4-dione |
|---|
| Traditional Name | 2-hydroxy-5-methoxy-3-[(8E,11E)-pentadeca-8,11,14-trien-1-yl]cyclohexa-2,5-diene-1,4-dione |
|---|
| SMILES | COC1=CC(=O)C(O)=C(CCCCCCC\C=C\C\C=C\CC=C)C1=O |
|---|
| InChI Identifier | InChI=1S/C22H30O4/c1-3-4-5-6-7-8-9-10-11-12-13-14-15-16-18-21(24)19(23)17-20(26-2)22(18)25/h3,5-6,8-9,17,24H,1,4,7,10-16H2,2H3/b6-5+,9-8+ |
|---|
| InChI Key | FGWRUVXUQWGLOX-HHWLVVFRSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as p-benzoquinones. These are benzoquinones where the two C=O groups are attached at the 1- and 4-positions, respectively. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbonyl compounds |
|---|
| Direct Parent | P-benzoquinones |
|---|
| Alternative Parents | |
|---|
| Substituents | - P-benzoquinone
- Vinylogous ester
- Vinylogous acid
- Enol
- Organic oxide
- Hydrocarbon derivative
- Aliphatic homomonocyclic compound
|
|---|
| Molecular Framework | Aliphatic homomonocyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00ej-3983000000-67e1459d4add48fcc62a | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-01ba-5963200000-3417f8f42d9782486fee | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4i-1019000000-adb5f934835d5fc1c4e6 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-052f-5798000000-44084a70c800f1d199a3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0ktf-8980000000-ed4d066f0b0475e40505 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0009000000-7f8273c59d66a4cf03eb | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-2129000000-fdb2a2f15b175edfe862 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0ac3-9132000000-71057cd8c26a9a35c3ef | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0009000000-ae9bc7f859382786f97c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0zfr-1918000000-75dae210cd4458209540 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0l02-5592000000-0ef0e763b89eddaca110 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-05i9-1191000000-7e8dd59f615c156af11a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-014l-9750000000-431b0445a2de77310d4c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-016u-9800000000-0757f10829a01fd1ef33 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0033443 |
|---|
| FooDB ID | FDB011481 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00000254 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 30776999 |
|---|
| ChEBI ID | 172587 |
|---|
| PubChem Compound ID | 14427830 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|