| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 23:26:14 UTC |
|---|
| Update Date | 2016-11-09 01:18:22 UTC |
|---|
| Accession Number | CHEM027007 |
|---|
| Identification |
|---|
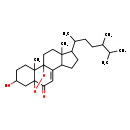
| Common Name | 5,9-Epidioxy-3-hydroxyergost-7-en-6-one |
|---|
| Class | Small Molecule |
|---|
| Description | 5,9-Epidioxy-3-hydroxyergost-7-en-6-one is found in mushrooms. 5,9-Epidioxy-3-hydroxyergost-7-en-6-one is a constituent of Hypsizygus marmoreus (bunashimeji) |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 5,9-Epidioxy-3-hydroxy-24-methylcholest-7-en-6-one | HMDB |
|
|---|
| Chemical Formula | C28H44O4 |
|---|
| Average Molecular Mass | 444.647 g/mol |
|---|
| Monoisotopic Mass | 444.324 g/mol |
|---|
| CAS Registry Number | 211486-17-8 |
|---|
| IUPAC Name | 8-(5,6-dimethylheptan-2-yl)-16-hydroxy-9,13-dimethyl-18,19-dioxapentacyclo[10.5.2.0¹,¹³.0⁴,¹².0⁵,⁹]nonadec-3-en-2-one |
|---|
| Traditional Name | 8-(5,6-dimethylheptan-2-yl)-16-hydroxy-9,13-dimethyl-18,19-dioxapentacyclo[10.5.2.0¹,¹³.0⁴,¹².0⁵,⁹]nonadec-3-en-2-one |
|---|
| SMILES | CC(C)C(C)CCC(C)C1CCC2C3=CC(=O)C45CC(O)CCC4(C)C3(CCC12C)OO5 |
|---|
| InChI Identifier | InChI=1S/C28H44O4/c1-17(2)18(3)7-8-19(4)21-9-10-22-23-15-24(30)28-16-20(29)11-12-26(28,6)27(23,31-32-28)14-13-25(21,22)5/h15,17-22,29H,7-14,16H2,1-6H3 |
|---|
| InChI Key | FZOPHVRKJMCTDV-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as ergostane steroids. These are steroids with a structure based on the ergostane skeleton, which arises from the methylation of cholestane at the 24-position. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Steroids and steroid derivatives |
|---|
| Sub Class | Ergostane steroids |
|---|
| Direct Parent | Ergostane steroids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Ergostane-skeleton
- Cyclohexenone
- Ortho-dioxolane
- Cyclic alcohol
- Dialkyl peroxide
- Secondary alcohol
- Ketone
- Oxacycle
- Organoheterocyclic compound
- Organic oxygen compound
- Organooxygen compound
- Alcohol
- Carbonyl group
- Hydrocarbon derivative
- Organic oxide
- Aliphatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aliphatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-016r-8013900000-37faff887579d0230be2 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-0udr-9411860000-ceec2ea1cec7ef798c69 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-004j-0001900000-793474d28471d8a92d01 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-002r-9226700000-57b2eb66765e9d158a2c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-9212000000-ad16460078172fdfd65b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0006-0000900000-06563854b897ee253664 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0006-0000900000-b5fcabdf719aa812bea7 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0f7a-2009300000-a9e46b13dc8332fd79a4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0002-0000900000-67dec7223cae3bfb2d94 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0aov-9205700000-c9f674cdc8ac6d9b666b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0536-9103100000-6c9296b80eca251c2066 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0006-0000900000-4a1a4016cf02e8b45f39 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0006-0000900000-4a1a4016cf02e8b45f39 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-0004900000-53e7b0fdf043b8b638d6 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0032667 |
|---|
| FooDB ID | FDB010620 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 74886385 |
|---|
| ChEBI ID | 173214 |
|---|
| PubChem Compound ID | 85218115 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|