| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 22:39:25 UTC |
|---|
| Update Date | 2016-11-09 01:18:10 UTC |
|---|
| Accession Number | CHEM025878 |
|---|
| Identification |
|---|
| Common Name | Glycosides |
|---|
| Class | Small Molecule |
|---|
| Description | Glycosides is found in allspice. Ouabain, a cardiac glycoside similar to digitoxin, is used to treat congestive heart failure and supraventricular arrhythmias due to reentry mechanisms, and to control ventricular rate in the treatment of chronic atrial fibrillation |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
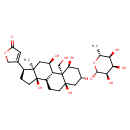
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| Glycoside | MeSH | | Kombetin | MeSH | | Strophosid | MeSH | | K-Strophanthin-beta | MeSH | | Acocantherin | HMDB | | Acolongifloriside K | HMDB | | Astrobain | HMDB | | g-Strophanthin | HMDB | | g-Strophicor | HMDB | | Gratibain | HMDB | | Gratus strophanthin | HMDB | | Ouabagenin L-rhamnoside | HMDB | | Ouabain | HMDB | | Ouabaine | HMDB | | Purostrophan | HMDB | | Rectobaina | HMDB | | Solufantina | HMDB | | Strodival | HMDB | | Strophalen | HMDB | | Strophanthin g | HMDB | | Strophoperm | HMDB | | Strophosan | HMDB | | Uabaina | HMDB | | Glycosides | MeSH | | Acolongifloroside K | MeSH | | g Strophanthin | MeSH |
|
|---|
| Chemical Formula | C29H44O12 |
|---|
| Average Molecular Mass | 584.653 g/mol |
|---|
| Monoisotopic Mass | 584.283 g/mol |
|---|
| CAS Registry Number | Not Available |
|---|
| IUPAC Name | 4-[(1S,2R,3S,5S,7R,10R,11R,14S,15R,17R)-3,7,11,17-tetrahydroxy-2-(hydroxymethyl)-15-methyl-5-{[(2R,3R,4R,5S,6R)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxy}tetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]-2,5-dihydrofuran-2-one |
|---|
| Traditional Name | 4-[(1S,2R,3S,5S,7R,10R,11R,14S,15R,17R)-3,7,11,17-tetrahydroxy-2-(hydroxymethyl)-15-methyl-5-{[(2R,3R,4R,5S,6R)-3,4,5-trihydroxy-6-methyloxan-2-yl]oxy}tetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]-5H-furan-2-one |
|---|
| SMILES | C[C@H]1O[C@@H](O[C@H]2C[C@H](O)[C@]3(CO)[C@H]4[C@H](O)C[C@]5(C)[C@@H](CC[C@@]5(O)[C@@H]4CC[C@@]3(O)C2)C2=CC(=O)OC2)[C@H](O)[C@H](O)[C@@H]1O |
|---|
| InChI Identifier | InChI=1S/C29H44O12/c1-13-22(34)23(35)24(36)25(40-13)41-15-8-19(32)28(12-30)21-17(3-5-27(28,37)9-15)29(38)6-4-16(14-7-20(33)39-11-14)26(29,2)10-18(21)31/h7,13,15-19,21-25,30-32,34-38H,3-6,8-12H2,1-2H3/t13-,15+,16+,17-,18-,19+,21-,22-,23-,24-,25+,26-,27-,28-,29-/m1/s1 |
|---|
| InChI Key | LPMXVESGRSUGHW-NZMIQZKWSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as cardenolide glycosides and derivatives. Cardenolide glycosides and derivatives are compounds containing a carbohydrate glycosidically bound to the cardenolide moiety. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Steroids and steroid derivatives |
|---|
| Sub Class | Steroid lactones |
|---|
| Direct Parent | Cardenolide glycosides and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Cardanolide-glycoside
- Steroidal glycoside
- 19-hydroxysteroid
- 1-hydroxysteroid
- 14-hydroxysteroid
- 5-hydroxysteroid
- Hydroxysteroid
- 11-hydroxysteroid
- 11-alpha-hydroxysteroid
- Hexose monosaccharide
- Glycosyl compound
- O-glycosyl compound
- 2-furanone
- Monosaccharide
- Oxane
- Tertiary alcohol
- Cyclic alcohol
- Dihydrofuran
- Enoate ester
- Alpha,beta-unsaturated carboxylic ester
- Secondary alcohol
- Carboxylic acid ester
- Lactone
- Polyol
- Acetal
- Carboxylic acid derivative
- Monocarboxylic acid or derivatives
- Oxacycle
- Organoheterocyclic compound
- Organic oxygen compound
- Organooxygen compound
- Alcohol
- Primary alcohol
- Organic oxide
- Hydrocarbon derivative
- Carbonyl group
- Aliphatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aliphatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0aou-0209050000-ff5d29b9a71cc0dd04e9 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-004u-2506409000-f6e6b4b9ec8f6ae25647 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS ("Glycosides,1TMS,#1" TMS) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_7) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_8) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_2) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_3) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_4) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_5) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_6) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_7) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_8) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_9) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_10) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_11) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_12) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_13) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_14) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_15) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00r2-0000490000-ba806aa277f179583eec | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00di-0101930000-7f40284ce7a78f730699 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00di-1112910000-f68efe23826a53a2145e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-015i-0000590000-b6bd31b640d6b6350b74 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-014r-1101940000-a2d91864bf2df81659a2 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4u-2005900000-252c65c11aeae5b683b8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-0000090000-968482c8635e1619bf2d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-00yi-5001790000-064fb537e519f8cf8882 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0ar9-9400830000-73dd15954b9792c10eff | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0000590000-2a960a4409b7608be7c5 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00ri-0015950000-de41dfd1a10d9666d965 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00du-9403810000-bf99265a71ab714ec04c | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0031448 |
|---|
| FooDB ID | FDB005482 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00003633 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Glycoside |
|---|
| Chemspider ID | 553182 |
|---|
| ChEBI ID | 24400 |
|---|
| PubChem Compound ID | 637579 |
|---|
| Kegg Compound ID | C01443 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Elliot GA: Unexpected natural death of iatrogenic origin. Forensic Sci. 1975 Feb;5(1):21-31. | | 2. Zaslavskaia RM, Logvinenko SI, Teiblium MM, Khalberg F: [Time organization of blood coagulation in patients with rheumatic heart disease involving circulation insufficiency]. Klin Med (Mosk). 1996;74(8):25-9. | | 3. De Tommasi N, Conti C, Stein ML, Pizza C: Structure and in vitro antiviral activity of triterpenoid saponins from Calendula arvensis. Planta Med. 1991 Jun;57(3):250-3. | | 4. Brower LP, Ryerson WN, Coppinger LL, Glazier SC: Ecological chemistry and the palatability spectrum. Science. 1968 Sep 27;161(3848):1349-50. | | 5. Takechi M, Uno C, Tanaka Y: Structure-activity relationships of synthetic saponins. Phytochemistry. 1996 Jan;41(1):121-3. | | 6. Simons K, Toomre D: Lipid rafts and signal transduction. Nat Rev Mol Cell Biol. 2000 Oct;1(1):31-9. | | 7. Watson AD: Thematic review series: systems biology approaches to metabolic and cardiovascular disorders. Lipidomics: a global approach to lipid analysis in biological systems. J Lipid Res. 2006 Oct;47(10):2101-11. Epub 2006 Aug 10. | | 8. Sethi JK, Vidal-Puig AJ: Thematic review series: adipocyte biology. Adipose tissue function and plasticity orchestrate nutritional adaptation. J Lipid Res. 2007 Jun;48(6):1253-62. Epub 2007 Mar 20. | | 9. Lingwood D, Simons K: Lipid rafts as a membrane-organizing principle. Science. 2010 Jan 1;327(5961):46-50. doi: 10.1126/science.1174621. | | 10. Duke, James A. (1992) Handbook of phytochemical constituents of GRAS herbs and other economic plants. Boca Raton, FL. CRC Press. | | 11. The lipid handbook with CD-ROM |
|
|---|