| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 21:18:12 UTC |
|---|
| Update Date | 2016-11-09 01:17:46 UTC |
|---|
| Accession Number | CHEM023802 |
|---|
| Identification |
|---|
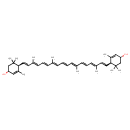
| Common Name | Lactucaxanthin |
|---|
| Class | Small Molecule |
|---|
| Description | A tunaxanthin that consists of epsilon,epsilon-carotene bearing hydroxy substituents at positions 3 and 3' (the 3S,3'S,6S,6'S-diastereomer). |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| (3S,3's,6S,6's)-Ε,ε-carotene-3,3'-diol | HMDB | | (3S,3's,6S,6's)-epsilon,epsilon-Carotene-3,3'-diol | HMDB | | Tunaxanthin F | HMDB | | Lactucaxanthin | SMPDB |
|
|---|
| Chemical Formula | C40H56O2 |
|---|
| Average Molecular Mass | 568.886 g/mol |
|---|
| Monoisotopic Mass | 568.428 g/mol |
|---|
| CAS Registry Number | 78306-12-4 |
|---|
| IUPAC Name | (1R,4R)-4-[(1E,3E,5E,7E,9E,11E,13E,15E,17E)-18-[(1R,4R)-4-hydroxy-2,6,6-trimethylcyclohex-2-en-1-yl]-3,7,12,16-tetramethyloctadeca-1,3,5,7,9,11,13,15,17-nonaen-1-yl]-3,5,5-trimethylcyclohex-2-en-1-ol |
|---|
| Traditional Name | lactucaxanthin |
|---|
| SMILES | C\C(\C=C\C=C(/C)\C=C\[C@H]1C(C)=C[C@H](O)CC1(C)C)=C/C=C/C=C(\C)/C=C/C=C(\C)/C=C/[C@H]1C(C)=C[C@H](O)CC1(C)C |
|---|
| InChI Identifier | InChI=1S/C40H56O2/c1-29(17-13-19-31(3)21-23-37-33(5)25-35(41)27-39(37,7)8)15-11-12-16-30(2)18-14-20-32(4)22-24-38-34(6)26-36(42)28-40(38,9)10/h11-26,35-38,41-42H,27-28H2,1-10H3/b12-11+,17-13+,18-14+,23-21+,24-22+,29-15+,30-16+,31-19+,32-20+/t35-,36-,37-,38-/m0/s1 |
|---|
| InChI Key | BIPAHAFBQLWRMC-SUOWZELTSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as xanthophylls. These are carotenoids containing an oxygenated carotene backbone. Carotenes are characterized by the presence of two end-groups (mostly cyclohexene rings, but also cyclopentene rings or acyclic groups) linked by a long branched alkyl chain. Carotenes belonging form a subgroup of the carotenoids family. Xanthophylls arise by oxygenation of the carotene backbone. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Tetraterpenoids |
|---|
| Direct Parent | Xanthophylls |
|---|
| Alternative Parents | |
|---|
| Substituents | - Xanthophyll
- Secondary alcohol
- Organic oxygen compound
- Hydrocarbon derivative
- Organooxygen compound
- Alcohol
- Aliphatic homomonocyclic compound
|
|---|
| Molecular Framework | Aliphatic homomonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-004i-3100029000-0177f8208fce473bb6aa | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0udi-3100190000-3b2c052d1ef9e5d4be0a | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS ("Lactucaxanthin,1TMS,#1" TMS) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_1) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0gb9-0101190000-5985ef12ae8dca4d83f2 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0frt-0719740000-cde5943fb9c3e4fe9f9e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0002-1449630000-1f7481e3eb567c876c85 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014i-0000090000-c3a0b350154f8c6bf618 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-014i-0000090000-fc5067576c09502ec0d5 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0udr-0522590000-a661bbd8ee97008407a6 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0fri-0131930000-1b4a9df72b953b7c578c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0f8c-0113920000-35ba818f2f9495497b11 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0wmr-0179400000-e432857c73a24a909aab | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014i-0202090000-e7ff503087b047287234 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-014i-0405490000-354e2648d408d044dc17 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-020a-0419300000-f98a6fa25c8795ec1019 | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0112256 |
|---|
| FooDB ID | FDB001534 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00003775 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 4444654 |
|---|
| ChEBI ID | 6357 |
|---|
| PubChem Compound ID | 5281242 |
|---|
| Kegg Compound ID | C08599 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. https://www.ncbi.nlm.nih.gov/pubmed/?term=19392686 | | 2. Khachik F, Spangler CJ, Smith JC Jr, Canfield LM, Steck A, Pfander H: Identification, quantification, and relative concentrations of carotenoids and their metabolites in human milk and serum. Anal Chem. 1997 May 15;69(10):1873-81. | | 3. Khachik F, Beecher GR, Goli MB, Lusby WR, Smith JC Jr: Separation and identification of carotenoids and their oxidation products in the extracts of human plasma. Anal Chem. 1992 Sep 15;64(18):2111-22. | | 4. Fraser PD, Bramley PM: The biosynthesis and nutritional uses of carotenoids. Prog Lipid Res. 2004 May;43(3):228-65. | | 5. Simons K, Toomre D: Lipid rafts and signal transduction. Nat Rev Mol Cell Biol. 2000 Oct;1(1):31-9. | | 6. Watson AD: Thematic review series: systems biology approaches to metabolic and cardiovascular disorders. Lipidomics: a global approach to lipid analysis in biological systems. J Lipid Res. 2006 Oct;47(10):2101-11. Epub 2006 Aug 10. | | 7. Sethi JK, Vidal-Puig AJ: Thematic review series: adipocyte biology. Adipose tissue function and plasticity orchestrate nutritional adaptation. J Lipid Res. 2007 Jun;48(6):1253-62. Epub 2007 Mar 20. | | 8. Lingwood D, Simons K: Lipid rafts as a membrane-organizing principle. Science. 2010 Jan 1;327(5961):46-50. doi: 10.1126/science.1174621. | | 9. The lipid handbook with CD-ROM |
|
|---|