| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 21:12:35 UTC |
|---|
| Update Date | 2016-11-09 01:17:45 UTC |
|---|
| Accession Number | CHEM023651 |
|---|
| Identification |
|---|
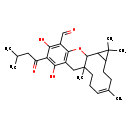
| Common Name | Euglobal VII |
|---|
| Class | Small Molecule |
|---|
| Description | Euglobal VII is a constituent of Eucalyptus globulus (Tasmanian blue gum) |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | Not Available |
|---|
| Chemical Formula | C28H38O5 |
|---|
| Average Molecular Mass | 454.598 g/mol |
|---|
| Monoisotopic Mass | 454.272 g/mol |
|---|
| CAS Registry Number | 77794-64-0 |
|---|
| IUPAC Name | (7Z)-14,16-dihydroxy-3,3,7,11-tetramethyl-15-(3-methylbutanoyl)-19-oxatetracyclo[9.8.0.0²,⁴.0¹³,¹⁸]nonadeca-7,13(18),14,16-tetraene-17-carbaldehyde |
|---|
| Traditional Name | (7Z)-14,16-dihydroxy-3,3,7,11-tetramethyl-15-(3-methylbutanoyl)-19-oxatetracyclo[9.8.0.0²,⁴.0¹³,¹⁸]nonadeca-7,13(18),14,16-tetraene-17-carbaldehyde |
|---|
| SMILES | CC(C)CC(=O)C1=C(O)C2=C(OC3C4C(CC\C(C)=C/CCC3(C)C2)C4(C)C)C(C=O)=C1O |
|---|
| InChI Identifier | InChI=1S/C28H38O5/c1-15(2)12-20(30)21-23(31)17-13-28(6)11-7-8-16(3)9-10-19-22(27(19,4)5)26(28)33-25(17)18(14-29)24(21)32/h8,14-15,19,22,26,31-32H,7,9-13H2,1-6H3/b16-8- |
|---|
| InChI Key | JIUCFHYHXVNZMU-PXNMLYILSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as germacrane sesquiterpenoids. These are sesquiterpenoids having the germacrane skeleton, with a structure characterized by a cyclodecane ring substituted with an isopropyl and two methyl groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Sesquiterpenoids |
|---|
| Direct Parent | Germacrane sesquiterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Germacrane sesquiterpenoid
- Butyrophenone
- Chromane
- Benzopyran
- 1-benzopyran
- Aryl ketone
- Aryl alkyl ketone
- Alkyl aryl ether
- Aryl-aldehyde
- Benzenoid
- Vinylogous acid
- Ketone
- Organoheterocyclic compound
- Oxacycle
- Ether
- Aldehyde
- Organooxygen compound
- Hydrocarbon derivative
- Organic oxide
- Organic oxygen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-01p6-5114900000-d8a245120dccdaf3b110 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-001l-4210090000-9f116ab674c360db978a | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4r-1092800000-9fea923e332d6597d64e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-052v-7595300000-15c28a5413a2000a93a7 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0aor-9380100000-d99be8939ca40339278a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0012900000-efcb9ee921c7eeef127c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0gb9-4039700000-1627736f3db60d0fe124 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-000j-9472200000-01c1ce1ea7a4382954d9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0000900000-0db53eae8610a0e47ec3 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0udi-0004900000-aa0a0952bede4a42d16e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9014300000-7977e8ee4be3498e9768 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4i-0000900000-76cc1ece83f9c07d0f80 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4i-0002900000-cb52b55a18453c38a640 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0pvm-2009000000-b373f02d1700a3c5a7ee | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0030037 |
|---|
| FooDB ID | FDB001339 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 35013121 |
|---|
| ChEBI ID | 175630 |
|---|
| PubChem Compound ID | 131750947 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | |
|---|