| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 20:49:44 UTC |
|---|
| Update Date | 2016-11-09 01:17:38 UTC |
|---|
| Accession Number | CHEM023095 |
|---|
| Identification |
|---|
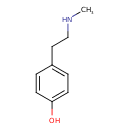
| Common Name | N-Methyltyramine |
|---|
| Class | Small Molecule |
|---|
| Description | |
|---|
| Contaminant Sources | |
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 4-Hydroxy-N-methylphenethylamine | ChEBI | | Methyl-4-tyramine | ChEBI | | p-(2-Methylaminoethyl)phenol | ChEBI | | 4-(2-(Methylamino)ethyl)-phenol | HMDB | | 4-(2-Methylaminoethyl)phenol | HMDB | | 4-[2-(Methylamino)ethyl]-phenol | HMDB | | 4-[2-(Methylamino)ethyl]phenol | HMDB | | Methyl tyramine | HMDB | | N-Methyl-p-tyramine | HMDB | | N-Methyltyraminium | HMDB | | NMT | HMDB | | p-(2-(Methylamino)ethyl)phenol | HMDB | | p-[2-(Methylamino)ethyl]-phenol | HMDB | | {p-[2-(methylamino)ethyl]phenol} | HMDB | | N-Methyltyrosamine | HMDB | | Methyl-4-tyramine hydrochloride | HMDB | | Methyl-4-tyramine hydrobromide | HMDB |
|
|---|
| Chemical Formula | C9H13NO |
|---|
| Average Molecular Mass | 151.206 g/mol |
|---|
| Monoisotopic Mass | 151.100 g/mol |
|---|
| CAS Registry Number | 370-98-9 |
|---|
| IUPAC Name | 4-[2-(methylamino)ethyl]phenol |
|---|
| Traditional Name | N-methyltyramine |
|---|
| SMILES | CNCCC1=CC=C(O)C=C1 |
|---|
| InChI Identifier | InChI=1S/C9H13NO/c1-10-7-6-8-2-4-9(11)5-3-8/h2-5,10-11H,6-7H2,1H3 |
|---|
| InChI Key | AXVZFRBSCNEKPQ-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as phenethylamines. Phenethylamines are compounds containing a phenethylamine moiety, which consists of a phenyl group substituted at the second position by an ethan-1-amine. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Benzene and substituted derivatives |
|---|
| Sub Class | Phenethylamines |
|---|
| Direct Parent | Phenethylamines |
|---|
| Alternative Parents | |
|---|
| Substituents | - Phenethylamine
- 1-hydroxy-2-unsubstituted benzenoid
- Aralkylamine
- Phenol
- Secondary amine
- Secondary aliphatic amine
- Organic nitrogen compound
- Organic oxygen compound
- Organopnictogen compound
- Hydrocarbon derivative
- Organooxygen compound
- Organonitrogen compound
- Amine
- Aromatic homomonocyclic compound
|
|---|
| Molecular Framework | Aromatic homomonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0006-9200000000-14293e75666be4e27019 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-00xr-7920000000-4709651ba3c22c23b463 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0uk9-0900000000-0f335d13104342fdeed5 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0fk9-2900000000-037595c7cc0c504058c6 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0kk9-9500000000-376efc98f806aac30929 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0900000000-27ea78b166d8b3172ce1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0udi-1900000000-fb693eca1267a0ba9685 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-05al-6900000000-6d964137e90c9e3293d1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00di-1900000000-6ac9e0419aa31060260b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00dl-7900000000-7fd4cbc25b6ff2a2bf1e | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-004l-9100000000-7021336d37b527e2e5fd | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0900000000-1c71ef4ed8c1b09fb863 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0udi-0900000000-4f21e6ca90b01b880bf4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-00kf-9600000000-4a0119f5c8223bec2ae4 | Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0006-9000000000-89ff5a43bc3e7737bcb6 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0003633 |
|---|
| FooDB ID | FDB000435 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | C00027432 |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | N-Methyltyramine |
|---|
| Chemspider ID | 9345 |
|---|
| ChEBI ID | 17458 |
|---|
| PubChem Compound ID | 9727 |
|---|
| Kegg Compound ID | C02442 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | M2MDB004568 |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Kimura T, Iwasaki N, Yokoe JI, Haruta S, Yokoo Y, Ogawara KI, Higaki K: Analysis and prediction of absorption profile including hepatic first-pass metabolism of N-methyltyramine, a potent stimulant of gastrin release present in beer, after oral ingestion in rats by gastrointestinal-transit-absorption model. Drug Metab Dispos. 2000 May;28(5):577-81. | | 2. Zhao XW, Li JX, Zhu ZR, Sun DQ, Liu SC: Anti-shock effects of synthetic effective compositions of fructus aurantii immaturus. Experimental study and clinical observation. Chin Med J (Engl). 1989 Feb;102(2):91-3. | | 3. Thiele I, Swainston N, Fleming RM, Hoppe A, Sahoo S, Aurich MK, Haraldsdottir H, Mo ML, Rolfsson O, Stobbe MD, Thorleifsson SG, Agren R, Bolling C, Bordel S, Chavali AK, Dobson P, Dunn WB, Endler L, Hala D, Hucka M, Hull D, Jameson D, Jamshidi N, Jonsson JJ, Juty N, Keating S, Nookaew I, Le Novere N, Malys N, Mazein A, Papin JA, Price ND, Selkov E Sr, Sigurdsson MI, Simeonidis E, Sonnenschein N, Smallbone K, Sorokin A, van Beek JH, Weichart D, Goryanin I, Nielsen J, Westerhoff HV, Kell DB, Mendes P, Palsson BO: A community-driven global reconstruction of human metabolism. Nat Biotechnol. 2013 May;31(5):419-25. doi: 10.1038/nbt.2488. Epub 2013 Mar 3. |
|
|---|