| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 18:54:15 UTC |
|---|
| Update Date | 2016-11-09 01:17:35 UTC |
|---|
| Accession Number | CHEM022804 |
|---|
| Identification |
|---|
| Common Name | Levoinositol |
|---|
| Class | Small Molecule |
|---|
| Description | |
|---|
| Contaminant Sources | - FooDB Chemicals
- HMDB Contaminants - Feces
|
|---|
| Contaminant Type | Not Available |
|---|
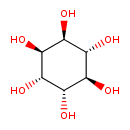
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| (-)-Inositol | ChEBI | | (1R,2R,3R,4R,5S,6S)-Cyclohexane-1,2,3,4,5,6-hexol | ChEBI | | 1,2,4/3,5,6-cyclohexanehexol | ChEBI | | L-Inositol | ChEBI |
|
|---|
| Chemical Formula | C6H12O6 |
|---|
| Average Molecular Mass | 180.156 g/mol |
|---|
| Monoisotopic Mass | 180.063 g/mol |
|---|
| CAS Registry Number | 551-72-4 |
|---|
| IUPAC Name | (1R,2R,3R,4R,5S,6S)-cyclohexane-1,2,3,4,5,6-hexol |
|---|
| Traditional Name | L-chiro-inositol |
|---|
| SMILES | [H][C@]1(O)[C@]([H])(O)[C@@]([H])(O)[C@]([H])(O)[C@]([H])(O)[C@@]1([H])O |
|---|
| InChI Identifier | InChI=1S/C6H12O6/c7-1-2(8)4(10)6(12)5(11)3(1)9/h1-12H/t1-,2-,3-,4-,5+,6+/m1/s1 |
|---|
| InChI Key | CDAISMWEOUEBRE-SHFUYGGZSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as cyclohexanols. Cyclohexanols are compounds containing an alcohol group attached to a cyclohexane ring. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Alcohols and polyols |
|---|
| Direct Parent | Cyclohexanols |
|---|
| Alternative Parents | |
|---|
| Substituents | - Cyclohexanol
- Sugar alcohol
- Cyclitol or derivatives
- Cyclic alcohol
- Polyol
- Hydrocarbon derivative
- Aliphatic homomonocyclic compound
|
|---|
| Molecular Framework | Aliphatic homomonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| GC-MS | GC-MS Spectrum - GC-MS (Non-derivatized) | splash10-014i-0359000000-cec32f7b73a366b3f067 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-03dr-9700000000-6cc8567e1b238f96f926 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-0900000000-5cb39f92646251665ebf | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-001i-0900000000-2aeb7708f31c772e2d95 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-08gi-8900000000-74f89711a1c6891808a4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-0900000000-8b68ff47f846a8ca1ddd | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-1900000000-03ef8572289795275d9c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0570-9300000000-950a37bd331028801e9f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-0900000000-c98742ed2c27fed8805b | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0570-9700000000-4e8df481dc1f1c124743 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-9000000000-ea5996664694b27554a8 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-0900000000-a3b1806f4d2d4bc9e470 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03e9-9500000000-40e1f34ad1ad2b894cd9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03dl-9000000000-5d72027b1e4acffd4c29 | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0034220 |
|---|
| FooDB ID | FDB001138 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 10199754 |
|---|
| ChEBI ID | 27374 |
|---|
| PubChem Compound ID | Not Available |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | M2MDB004665 |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Dai X, Zhang L, Hong T: Host cellular signaling induced by influenza virus. Sci China Life Sci. 2011 Jan;54(1):68-74. doi: 10.1007/s11427-010-4116-z. Epub 2011 Jan 21. | | 2. Cadman ED, Naugles DD, Lee CM: cAMP is not involved in interleukin-1-induced interleukin-6 release from human astrocytoma cells. Neurosci Lett. 1994 Sep 12;178(2):251-4. | | 3. Chen G, Kirby M, Zeng W, Young NS, Maciejewski JP: Superior growth of glycophosphatidy linositol-anchored protein-deficient progenitor cells in vitro is due to the higher apoptotic rate of progenitors with normal phenotype in vivo. Exp Hematol. 2002 Jul;30(7):774-82. | | 4. Kim CH, Park YS, Chung KN, Elwood PC: Sorting of the human folate receptor in MDCK cells. J Biochem Mol Biol. 2004 May 31;37(3):362-9. | | 5. Butlen D, Bernard C, Ammar A, Ferrary E: Purine and pyrimidine nucleotide-sensitive phosphoinositidase C in ampulla from frog semicircular canal. Am J Physiol. 1997 Jan;272(1 Pt 2):R51-8. | | 6. Pak Y, Huang LC, Lilley KJ, Larner J: In vivo conversion of [3H]myoinositol to [3H]chiroinositol in rat tissues. J Biol Chem. 1992 Aug 25;267(24):16904-10. | | 7. Matsumoto T, Kawanobe Y, Morita K, Ogata E: Effect of 1,25-dihydroxyvitamin D3 on phospholipid metabolism in a clonal osteoblast-like rat osteogenic sarcoma cell line. J Biol Chem. 1985 Nov 5;260(25):13704-9. | | 8. Takeuchi K, Shibamoto S, Nagamine K, Shigemori I, Omura S, Kitamura N, Ito F: Signaling pathways leading to transcription and translation cooperatively regulate the transient increase in expression of c-Fos protein. J Biol Chem. 2001 Jul 13;276(28):26077-83. Epub 2001 May 14. | | 9. Bates SH, Jones RB, Bailey CJ: Insulin-like effect of pinitol. Br J Pharmacol. 2000 Aug;130(8):1944-8. | | 10. Warita H, Manabe Y, Murakami T, Shiro Y, Nagano I, Abe K: Early decrease of survival signal-related proteins in spinal motor neurons of presymptomatic transgenic mice with a mutant SOD1 gene. Apoptosis. 2001 Oct;6(5):345-52. | | 11. Pampillo M, Camuso N, Taylor JE, Szereszewski JM, Ahow MR, Zajac M, Millar RP, Bhattacharya M, Babwah AV: Regulation of GPR54 signaling by GRK2 and {beta}-arrestin. Mol Endocrinol. 2009 Dec;23(12):2060-74. doi: 10.1210/me.2009-0013. Epub 2009 Oct 21. | | 12. Anzai-Takeda Y, Takeda Y, Sendo F, Araki Y: Inhibition of cell spreading in CHO cells transfected with cDNA of a glycosylphosphatidyl inositol-anchored protein, GPI-80. Immunobiology. 2005;210(1):1-10. | | 13. Pak Y, Larner J: Identification and characterization of chiroinositol-containing phospholipids from bovine liver. Biochem Biophys Res Commun. 1992 Apr 30;184(2):1042-7. | | 14. Yannai, Shmuel. (2004) Dictionary of food compounds with CD-ROM: Additives, flavors, and ingredients. Boca Raton: Chapman & Hall/CRC. |
|
|---|