| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 18:18:56 UTC |
|---|
| Update Date | 2016-11-09 01:17:25 UTC |
|---|
| Accession Number | CHEM022064 |
|---|
| Identification |
|---|
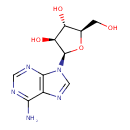
| Common Name | Vidarabine |
|---|
| Class | Small Molecule |
|---|
| Description | A purine nucleoside in which adenine is attached to arabinofuranose via a beta-N(9)-glycosidic bond. |
|---|
| Contaminant Sources | - HMDB Contaminants - Urine
- STOFF IDENT Compounds
|
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 2-(6-AMINO-purin-9-yl)-5-hydroxymethyl-tetrahydro-furan-3,4-diol | ChEBI | | 9-beta-D-Arabinofuranosyl-9H-purin-6-amine | ChEBI | | 9-beta-D-Arabinofuranosyladenine | ChEBI | | Spongoadenosine | ChEBI | | Vidarabine anhydrous | Kegg | | ARA-a | Kegg | | Armes | Kegg | | 9-b-D-Arabinofuranosyl-9H-purin-6-amine | Generator | | 9-Β-D-arabinofuranosyl-9H-purin-6-amine | Generator | | 9-b-D-Arabinofuranosyladenine | Generator | | 9-Β-D-arabinofuranosyladenine | Generator | | 9-beta-D-Arabinofuranosyl-adenine | HMDB | | Adenine arabinoside | HMDB | | Araadenosine | HMDB | | Arabinoside adenine | HMDB | | Arabinosyl adenine | HMDB | | Arabinosyladenine | HMDB | | a, alpha-Ara | HMDB | | Ara a | HMDB | | Arabinofuranosyladenine | HMDB | | Monarch brand OF vidarabine | HMDB | | ViraA | HMDB | | alpha Ara a | HMDB | | alpha D Arabinofuranosyladenine | HMDB | | alpha-Ara a | HMDB | | 9 beta Arabinofuranosyladenine | HMDB | | 9 beta D Arabinofuranosyladenine | HMDB | | a, Ara | HMDB | | beta Ara a | HMDB | | Arabinoside, adenine | HMDB | | alpha-D-Arabinofuranosyladenine | HMDB | | 9-beta-Arabinofuranosyladenine | HMDB | | a, beta-Ara | HMDB | | Parke davis brand OF vidarabine | HMDB | | Vira a | HMDB | | Vira-a | HMDB | | beta-Ara a | HMDB | | Vidarabine | ChEBI |
|
|---|
| Chemical Formula | C10H13N5O4 |
|---|
| Average Molecular Mass | 267.241 g/mol |
|---|
| Monoisotopic Mass | 267.097 g/mol |
|---|
| CAS Registry Number | 24356-66-9 |
|---|
| IUPAC Name | (2R,3S,4S,5R)-2-(6-amino-9H-purin-9-yl)-5-(hydroxymethyl)oxolane-3,4-diol |
|---|
| Traditional Name | armes |
|---|
| SMILES | NC1=NC=NC2=C1N=CN2[C@@H]1O[C@H](CO)[C@@H](O)[C@@H]1O |
|---|
| InChI Identifier | InChI=1S/C10H13N5O4/c11-8-5-9(13-2-12-8)15(3-14-5)10-7(18)6(17)4(1-16)19-10/h2-4,6-7,10,16-18H,1H2,(H2,11,12,13)/t4-,6-,7+,10-/m1/s1 |
|---|
| InChI Key | OIRDTQYFTABQOQ-UHTZMRCNSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as purine nucleosides. Purine nucleosides are compounds comprising a purine base attached to a ribosyl or deoxyribosyl moiety. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Nucleosides, nucleotides, and analogues |
|---|
| Class | Purine nucleosides |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Purine nucleosides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Purine nucleoside
- Glycosyl compound
- N-glycosyl compound
- 6-aminopurine
- Pentose monosaccharide
- Imidazopyrimidine
- Purine
- Aminopyrimidine
- Monosaccharide
- N-substituted imidazole
- Pyrimidine
- Imidolactam
- Tetrahydrofuran
- Azole
- Imidazole
- Heteroaromatic compound
- Secondary alcohol
- Organoheterocyclic compound
- Azacycle
- Oxacycle
- Organic oxygen compound
- Organic nitrogen compound
- Alcohol
- Organonitrogen compound
- Hydrocarbon derivative
- Organopnictogen compound
- Organooxygen compound
- Amine
- Primary alcohol
- Primary amine
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0a4l-9350000000-ed8639e5b54e46825628 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (3 TMS) - 70eV, Positive | splash10-014i-1492400000-1fe7f0d7ca11fdb4800f | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-000i-2910000000-e18a8d00eb0e73aa6dec | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-000i-2910000000-e18a8d00eb0e73aa6dec | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0940000000-42bee9785f9b55b32eea | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0900000000-c8cda6a661ace060572c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-2900000000-0304a85c3951aa1cb41a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0159-0690000000-0258e5851611884483d9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-0900000000-e6b8fc6980387f646d64 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-053r-2900000000-1e0cad8d044715d90f12 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-0910000000-83e24e24fe97518d665a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-0900000000-291c06f7d909185dce29 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-053r-1900000000-1401974aef47ea042a5c | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0920000000-c89adfa0ec9ccc01f131 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0900000000-c1ee2199364d0258bae1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-1900000000-fd986996dd15b2469008 | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00194 |
|---|
| HMDB ID | HMDB0014340 |
|---|
| FooDB ID | Not Available |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | RAB |
|---|
| Wikipedia Link | Vidarabine |
|---|
| Chemspider ID | 20400 |
|---|
| ChEBI ID | 45327 |
|---|
| PubChem Compound ID | 21704 |
|---|
| Kegg Compound ID | C07195 |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Feher Z, Mishra NC: An aphidicolin-resistant mutant of Chinese hamster ovary cell with altered DNA polymerase and 3' exonuclease activities. Biochim Biophys Acta. 1995 Aug 22;1263(2):141-6. | | 2. Russell RR 3rd, Bergeron R, Shulman GI, Young LH: Translocation of myocardial GLUT-4 and increased glucose uptake through activation of AMPK by AICAR. Am J Physiol. 1999 Aug;277(2 Pt 2):H643-9. | | 3. Moore CL, Chiaramonte M, Higgins T, Kuchta RD: Synthesis of nucleotide analogues that potently and selectively inhibit human DNA primase. Biochemistry. 2002 Nov 26;41(47):14066-75. | | 4. Kuchta RD, Ilsley D, Kravig KD, Schubert S, Harris B: Inhibition of DNA primase and polymerase alpha by arabinofuranosylnucleoside triphosphates and related compounds. Biochemistry. 1992 May 19;31(19):4720-8. | | 5. Chen J, Hudson E, Chi MM, Chang AS, Moley KH, Hardie DG, Downs SM: AMPK regulation of mouse oocyte meiotic resumption in vitro. Dev Biol. 2006 Mar 15;291(2):227-38. Epub 2006 Jan 26. | | 6. Thompson HC, Kuchta RD: Arabinofuranosyl nucleotides are not chain-terminators during initiation of new strands of DNA by DNA polymerase alpha-primase. Biochemistry. 1995 Sep 5;34(35):11198-203. | | 7. Feher Z, Mishra NC: Aphidicolin-resistant Chinese hamster ovary cells possess altered DNA polymerases of the alpha-family. Biochim Biophys Acta. 1994 May 17;1218(1):35-47. |
|
|---|