| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 18:13:56 UTC |
|---|
| Update Date | 2016-11-09 01:17:22 UTC |
|---|
| Accession Number | CHEM021848 |
|---|
| Identification |
|---|
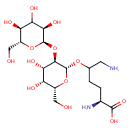
| Common Name | Glucosylgalactosyl hydroxylysine |
|---|
| Class | Small Molecule |
|---|
| Description | Glucosylgalactosyl hydroxylysine is a Glycoside of hydroxylysine. Hydroxylation of lysine is an important post-translational modification of collagen, (PMID 10928217) a reaction catalyzed by the enzyme Lysyl hydroxylase (EC 1.14.11.4) forming hydroxylysine in collagens and other proteins with collagen-like amino acid sequences, by the hydroxylation of lysine residues in X-lys-gly sequences.
The hydroxylysine residues formed in the lysyl hydroxylase reaction have 2 important functions: first, their hydroxy groups serve as sites of attachment for carbohydrate units, either the monosaccharide galactose or the disaccharide glucosylgalactose; and second, they are essential for the stability of the intermolecular collagen crosslinks. (OMIM 153454)
Hydroxylysine-deficient skin collagen is manifested in Ehlers-Danlos syndrome, type VIA, A heritable disorder of connective tissue (PMID 5016372) [HMDB] |
|---|
| Contaminant Sources | - FooDB Chemicals
- HMDB Contaminants - Urine
|
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 1,2alpha-Glucosylgalactosyl-O-hydroxylysine | HMDB | | 2-O-a-D-Glucopyranosyl-O-b-D-galactopyranosylhydroxylysine | HMDB | | 2-O-alpha-D-Glucopyranosyl-O-beta-D-galactopyranosylhydroxylysine | HMDB | | 2-O-alpha-delta-Glucopyranosyl-O-beta-delta-galactopyranosylhydroxylysine | HMDB | | 5-[O-a-D-Glucopyranosyl-(1->2)-b-D-galactopyranosyloxy]-L-lysine | HMDB | | 5-[O-alpha-D-Glucopyranosyl-(1->2)-beta-D-galactopyranosyloxy]-L-lysine | HMDB | | 5-[O-alpha-delta-Glucopyranosyl-(1->2)-beta-delta-galactopyranosyloxy]-L-lysine | HMDB | | Glucopyranosylgalactopyranosylhydroxylysine | HMDB | | Glucosido-galactosyl-hydroxylysine | HMDB | | Glucosyl galactosyl-D-hydroxylysine | HMDB | | Glucosyl galactosyl-delta-hydroxylysine | HMDB | | Glucosylgalactosylhydroxylysine | HMDB | | Hydroxylysine-galactose-glucose | HMDB | | Hydroxylysine-glucose-galactose | HMDB | | L-5-((2-O-alpha-D-Glucopyranosyl-beta-D-galactopyranosyl)oxy)-lysine | HMDB | | L-5-((2-O-alpha-delta-Glucopyranosyl-beta-delta-galactopyranosyl)oxy)-lysine | HMDB | | 1,2 alpha-Glucosylgalactosyl-O-hydroxylysine | HMDB | | (2S)-2,6-Diamino-5-{[(2R,3R,4S,5R,6R)-4,5-dihydroxy-6-(hydroxymethyl)-3-{[(2R,3R,5S,6R)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}oxan-2-yl]oxy}hexanoate | HMDB |
|
|---|
| Chemical Formula | C18H34N2O13 |
|---|
| Average Molecular Mass | 486.468 g/mol |
|---|
| Monoisotopic Mass | 486.206 g/mol |
|---|
| CAS Registry Number | 32448-35-4 |
|---|
| IUPAC Name | (2S)-2,6-diamino-5-{[(2R,3R,4S,5R,6R)-4,5-dihydroxy-6-(hydroxymethyl)-3-{[(2R,3R,5S,6R)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}oxan-2-yl]oxy}hexanoic acid |
|---|
| Traditional Name | (2S)-2,6-diamino-5-{[(2R,3R,4S,5R,6R)-4,5-dihydroxy-6-(hydroxymethyl)-3-{[(2R,3R,5S,6R)-3,4,5-trihydroxy-6-(hydroxymethyl)oxan-2-yl]oxy}oxan-2-yl]oxy}hexanoic acid |
|---|
| SMILES | NCC(CC[C@H](N)C(O)=O)O[C@@H]1O[C@H](CO)[C@H](O)[C@H](O)[C@H]1O[C@H]1O[C@H](CO)[C@@H](O)C(O)[C@H]1O |
|---|
| InChI Identifier | InChI=1S/C18H34N2O13/c19-3-6(1-2-7(20)16(28)29)30-18-15(13(26)11(24)9(5-22)32-18)33-17-14(27)12(25)10(23)8(4-21)31-17/h6-15,17-18,21-27H,1-5,19-20H2,(H,28,29)/t6?,7-,8+,9+,10+,11-,12?,13-,14+,15+,17+,18+/m0/s1 |
|---|
| InChI Key | UTIRJVJBKWSIOX-SRMFCGEKSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as fatty acyl glycosides of mono- and disaccharides. Fatty acyl glycosides of mono- and disaccharides are compounds composed of a mono- or disaccharide moiety linked to one hydroxyl group of a fatty alcohol or of a phosphorylated alcohol (phosphoprenols), a hydroxy fatty acid or to one carboxyl group of a fatty acid (ester linkage) or to an amino alcohol. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Fatty acyl glycosides |
|---|
| Direct Parent | Fatty acyl glycosides of mono- and disaccharides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Fatty acyl glycoside of mono- or disaccharide
- Disaccharide
- Glycosyl compound
- O-glycosyl compound
- Alpha-amino acid
- Alpha-amino acid or derivatives
- L-alpha-amino acid
- Medium-chain fatty acid
- Amino fatty acid
- Heterocyclic fatty acid
- Hydroxy fatty acid
- Fatty acid
- Oxane
- Amino acid or derivatives
- Amino acid
- Secondary alcohol
- Acetal
- Carboxylic acid derivative
- Carboxylic acid
- Oxacycle
- Organoheterocyclic compound
- Monocarboxylic acid or derivatives
- Polyol
- Organic oxygen compound
- Organic nitrogen compound
- Hydrocarbon derivative
- Alcohol
- Carbonyl group
- Organic oxide
- Primary amine
- Primary alcohol
- Primary aliphatic amine
- Amine
- Organopnictogen compound
- Organonitrogen compound
- Organooxygen compound
- Aliphatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aliphatic heteromonocyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-05q9-6125900000-0de572c676dd4f3e5e71 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-0hh9-9500134000-e2f856e837ca6a15e9b8 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_69) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_4_31) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_4_101) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_4_171) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_4_172) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_4_173) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_4_200) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_4_215) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_1) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_71) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_72) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_73) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_184) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_186) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_297) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_298) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_301) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_307) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_308) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_327) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_342) - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_5_349) - 70eV, Positive | Not Available | Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - CE-QTOF-MS system (Agilent 7100 CE + 6550 QTOF) 30V, Positive | splash10-03fs-0910000000-79c09465fc3edc81cabe | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-07bb-0924600000-194b269d426da482f1f9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-02bb-0922100000-f940966d1894b78b8a95 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03xs-1911000000-85b476e6f54ee25e2bb2 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-08g3-1904500000-c5212557085785075a62 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-03di-0901100000-cb344df736c38c1e895d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-03di-3900000000-c93fab31d1d97345d410 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0102900000-35cae597085109df5d12 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0cdu-6903500000-ca992383cfddb3d00925 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-05fu-9222000000-d094ca3b799998628fbf | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udr-0000900000-726c99d526e648fbee4d | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-004s-1902800000-7c98f8d7a23c2ffd7139 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-9202100000-4d1e13c37067063c24ec | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0000585 |
|---|
| FooDB ID | FDB022130 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | 5567 |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 109059 |
|---|
| ChEBI ID | 89573 |
|---|
| PubChem Compound ID | 122304 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Garza, Hector; Bennett, Nelson Jr.; Rodriguez, Gladys P. Improved rapid method for the isolation, purification and identification of collagen glycosides. Journal of Chromatography, A (1996), 732(2), 385-389. | | 2. Savolainen ER, Kero M, Pihlajaniemi T, Kivirikko KI: Deficiency of galactosylhydroxylysyl glucosyltransferase, an enzyme of collagen synthesis, in a family with dominant epidermolysis bullosa simplex. N Engl J Med. 1981 Jan 22;304(4):197-204. | | 3. Kelleher PC: Urinary excretion of hydroxyproline, hydroxylysine and hydroxylysine glycosides by patients with Paget's disease of bone and carcinoma with metastases in bone. Clin Chim Acta. 1979 Mar 15;92(3):373-9. | | 4. Schroder CH, Monnens LA, van Lith-Zanders HM, Trijbels JM, Veerkamp JH: The urinary excretion of total hydroxylysine and its glycosides in normal persons, and in patients suffering from Alport's syndrome--contribution of the peptide-bound fraction. Nephron. 1987;47(4):253-7. | | 5. Schroder CH, Langeveld JP, van Raay-Selten BH, Trijbels FJ, de Graaf R, Veerkamp JH, Monnens LA: Urinary excretion of hydroxylysine and its glycosides in normal persons of different ages--influence of maturation. Int J Pediatr Nephrol. 1985 Oct-Dec;6(4):239-44. | | 6. Schroder CH, Monnens LA, van Lith-Zanders HM, Trijbels JM, Veerkamp JH, Langeveld JP: Urinary excretion of hydroxylysine and its glycosides in Alport's syndrome and several other glomerulopathies. Nephron. 1986;44(2):103-7. | | 7. Grazioli V, Alfano M, Stenico A, Casari E: Urinary output of hydroxylysine glycosides and pyridinium cross-links in detecting rat bone collagen turnover rate. FEBS Lett. 1996 Jun 17;388(2-3):134-8. | | 8. Ono S, Shimizu N, Imai T, Rodriguez GP: Urinary collagen metabolite excretion in amyotrophic lateral sclerosis. Muscle Nerve. 2001 Jun;24(6):821-5. | | 9. Rodriguez GP, Claus-Walker J: Measurement of hydroxylysine glycosides in urine and its application to spinal cord injury. J Chromatogr. 1984 Jun 8;308:65-73. | | 10. Szulc P, Seeman E, Delmas PD: Biochemical measurements of bone turnover in children and adolescents. Osteoporos Int. 2000;11(4):281-94. | | 11. Pinnell SR, Krane SM, Kenzora JE, Glimcher MJ: A heritable disorder of connective tissue. Hydroxylysine-deficient collagen disease. N Engl J Med. 1972 May 11;286(19):1013-20. | | 12. Bank, Ruud A., Beekman, B., Tenni, R., TeKoppele, Johan M. Pre-column derivatization method for the measurement of glycosylated hydroxylysines of collagenous proteins. Journal of Chromatography, B: Biomedical Sciences and Applications (1997), 703(1 + 2), 267-272 | | 13. Kakimoto, Y., Akazawa, S. Isolation and identification of NG,NG- and NG,N'G-dimethylarginine, N -mono-, di-, and trimethyllysine, and glucosylgalactosyl- and galactosyl- -hydroxylysine from human urine. Journal of Biological Chemistry (1970), 245(21), 5751-8 |
|
|---|