| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-25 18:12:08 UTC |
|---|
| Update Date | 2016-11-09 01:17:21 UTC |
|---|
| Accession Number | CHEM021772 |
|---|
| Identification |
|---|
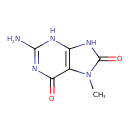
| Common Name | 8-Hydroxy-7-methylguanine |
|---|
| Class | Small Molecule |
|---|
| Description | An oxopurine that is guanine with an oxo group at position 8 and a methyl substituent at position 7. |
|---|
| Contaminant Sources | - FooDB Chemicals
- HMDB Contaminants - Urine
|
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | Not Available |
|---|
| Chemical Formula | C6H7N5O2 |
|---|
| Average Molecular Mass | 181.152 g/mol |
|---|
| Monoisotopic Mass | 181.060 g/mol |
|---|
| CAS Registry Number | 1688-85-3 |
|---|
| IUPAC Name | 2-amino-7-methyl-6,7,8,9-tetrahydro-3H-purine-6,8-dione |
|---|
| Traditional Name | 2-amino-7-methyl-3,9-dihydropurine-6,8-dione |
|---|
| SMILES | CN1C(=O)NC2=C1C(=O)N=C(N)N2 |
|---|
| InChI Identifier | InChI=1S/C6H7N5O2/c1-11-2-3(9-6(11)13)8-5(7)10-4(2)12/h1H3,(H4,7,8,9,10,12,13) |
|---|
| InChI Key | VHPXSVXJBWZORQ-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as hypoxanthines. Hypoxanthines are compounds containing the purine derivative 1H-purin-6(9H)-one. Purine is a bicyclic aromatic compound made up of a pyrimidine ring fused to an imidazole ring. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Imidazopyrimidines |
|---|
| Sub Class | Purines and purine derivatives |
|---|
| Direct Parent | Hypoxanthines |
|---|
| Alternative Parents | |
|---|
| Substituents | - 6-oxopurine
- Hypoxanthine
- Aminopyrimidine
- Pyrimidone
- N-substituted imidazole
- Pyrimidine
- Vinylogous amide
- Imidazole
- Azole
- Heteroaromatic compound
- Urea
- Azacycle
- Organic oxide
- Amine
- Primary amine
- Organooxygen compound
- Organonitrogen compound
- Organic nitrogen compound
- Organopnictogen compound
- Organic oxygen compound
- Hydrocarbon derivative
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-001r-0900000000-4e0a1211082933a52189 | Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-0900000000-7c1b5b61581c8650b910 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-001i-0900000000-15a7e1a0805e52238283 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00rb-7900000000-b9f171dba84ae611184a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-0900000000-ca5228f6b8181c8393ab | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001r-1900000000-8ad42ac014332bbefdc5 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0007-9200000000-a717cec7a8c28103e4aa | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-0900000000-90d288c41daa0d7f9fe7 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-0900000000-27306b4f0abc07f63593 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9200000000-0f5e5385863ab5177669 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-0900000000-e9758175f80068d624c4 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-001i-0900000000-0b6d69fb729f5067e205 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000l-5900000000-32125fca442ba581cefa | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0006037 |
|---|
| FooDB ID | FDB023813 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 272336 |
|---|
| ChEBI ID | 74064 |
|---|
| PubChem Compound ID | 308075 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Morris GS, Simmonds HA, Davies PM: Use of biological fluids for the rapid diagnosis of potentially lethal inherited disorders of human purine and pyrimidine metabolism. Biomed Chromatogr. 1986 Jun;1(3):109-18. | | 2. Litwack MD, Weissmann B: Source of urinary 8-hydroxy-7-methylguanine in man. Biochemistry. 1966 Sep;5(9):3007-12. | | 3. Choi JY, Guengerich FP: Kinetic evidence for inefficient and error-prone bypass across bulky N2-guanine DNA adducts by human DNA polymerase iota. J Biol Chem. 2006 May 5;281(18):12315-24. Epub 2006 Mar 8. | | 4. Choi JY, Guengerich FP: Adduct size limits efficient and error-free bypass across bulky N2-guanine DNA lesions by human DNA polymerase eta. J Mol Biol. 2005 Sep 9;352(1):72-90. | | 5. Porcelli B, Pagani R, Lorenzini L, De Martino A, Catinella S, Traldi P: Different mass spectrometric approaches in the identification of endogenous methylated purine bases in urine extracts. Rapid Commun Mass Spectrom. 1994 Jun;8(6):443-50. | | 6. Sander G, Topp H, Heller-Schoch G, Wieland J, Schoch G: Ribonucleic acid turnover in man:RNA catabolites in urine as measure for the metabolism of each of the three major species of RNA. Clin Sci (Lond). 1986 Oct;71(4):367-74. | | 7. Kanduc D, Sapia G: Origin of 1,7-dimethylguanosine in tRNA chemical and enzymatic methylation. Boll Soc Ital Biol Sper. 1982 Oct 15;58(19):1221-5. | | 8. Niwa T, Takeda N, Yoshizumi H: RNA metabolism in uremic patients: accumulation of modified ribonucleosides in uremic serum. Technical note. Kidney Int. 1998 Jun;53(6):1801-6. |
|
|---|