| Record Information |
|---|
| Version | 1.0 |
|---|
| Creation Date | 2016-05-19 03:49:29 UTC |
|---|
| Update Date | 2016-11-09 01:14:30 UTC |
|---|
| Accession Number | CHEM012086 |
|---|
| Identification |
|---|
| Common Name | Mannanase, endo-1,4-.beta.- |
|---|
| Class | Small Molecule |
|---|
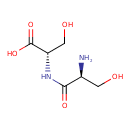
| Description | A dipeptide formed from two L-serine residues. |
|---|
| Contaminant Sources | - HPV EPA Chemicals
- STOFF IDENT Compounds
|
|---|
| Contaminant Type | Not Available |
|---|
| Chemical Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| H-L-Ser-L-ser-OH | ChEBI | | H-Ser-ser-OH | ChEBI | | L-Ser-L-ser | ChEBI | | S-S | ChEBI | | SS | ChEBI | | (2S)-2-{[(2S)-2-amino-1,3-dihydroxypropylidene]amino}-3-hydroxypropanoate | HMDB | | L-Seryl-L-serine | HMDB | | N-L-Seryl-L-serine | HMDB | | N-Serylserine | HMDB | | S-S Dipeptide | HMDB | | SS Dipeptide | HMDB | | Ser-ser | HMDB | | Serine serine dipeptide | HMDB | | Serine-serine dipeptide | HMDB | | Serinyl-serine | HMDB | | Serinylserine | HMDB | | Seryl-serine | HMDB | | Serylserine | ChEBI |
|
|---|
| Chemical Formula | C6H12N2O5 |
|---|
| Average Molecular Mass | 192.171 g/mol |
|---|
| Monoisotopic Mass | 192.075 g/mol |
|---|
| CAS Registry Number | 37288-54-3 |
|---|
| IUPAC Name | (2S)-2-[(2S)-2-amino-3-hydroxypropanamido]-3-hydroxypropanoic acid |
|---|
| Traditional Name | L-serine, N-L-seryl- |
|---|
| SMILES | [H][C@](N)(CO)C(O)=N[C@@]([H])(CO)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C6H12N2O5/c7-3(1-9)5(11)8-4(2-10)6(12)13/h3-4,9-10H,1-2,7H2,(H,8,11)(H,12,13)/t3-,4-/m0/s1 |
|---|
| InChI Key | XZKQVQKUZMAADP-IMJSIDKUSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as dipeptides. These are organic compounds containing a sequence of exactly two alpha-amino acids joined by a peptide bond. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | Dipeptides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Alpha-dipeptide
- N-acyl-l-alpha-amino acid
- N-acyl-alpha-amino acid
- N-acyl-alpha amino acid or derivatives
- Alpha-amino acid amide
- Serine or derivatives
- Alpha-amino acid or derivatives
- Beta-hydroxy acid
- Hydroxy acid
- Amino acid or derivatives
- Amino acid
- Carboxamide group
- Carboxylic acid salt
- Secondary carboxylic acid amide
- Carboxylic acid
- Monocarboxylic acid or derivatives
- Alcohol
- Organonitrogen compound
- Organooxygen compound
- Primary alcohol
- Primary aliphatic amine
- Primary amine
- Organic zwitterion
- Organic salt
- Hydrocarbon derivative
- Carbonyl group
- Organic oxide
- Organopnictogen compound
- Amine
- Organic oxygen compound
- Organic nitrogen compound
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | Not Available |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Not Available |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-01tc-5900000000-a45243c43945dbd2e982 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03di-9400000000-55bb53f88f5b99210438 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0006-9000000000-d41d6623979c159ec935 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-01ox-0900000000-cb44a137581c072567ef | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-03mv-2900000000-b2f2608daed8f477aa68 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4l-9100000000-f5e94186198db822996f | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0btc-4900000000-319f81bc5a65d28953a1 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03di-9100000000-0b2579ddaf96b34f62b9 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-03dl-9000000000-d8c29fe92e027a3c30cd | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-006x-2900000000-7beeec871e2162a3852a | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-00di-9200000000-4782d9a93fe0d28ab3f6 | Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0596-9000000000-46e7e9ec9081b97500bc | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum | | 1D NMR | 13C NMR Spectrum | Not Available | Spectrum | | 1D NMR | 1H NMR Spectrum | Not Available | Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not Available |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB0029048 |
|---|
| FooDB ID | FDB112055 |
|---|
| Phenol Explorer ID | Not Available |
|---|
| KNApSAcK ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| Chemspider ID | 5382072 |
|---|
| ChEBI ID | 73653 |
|---|
| PubChem Compound ID | 7019104 |
|---|
| Kegg Compound ID | Not Available |
|---|
| YMDB ID | Not Available |
|---|
| ECMDB ID | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | | 1. Golimbet VE, Lebedeva IS, Monakhov MV, Korovaitseva GI, Lezheiko TV, Abramova LI, Kaleda VG, Karpov VL: [The Cys allele of the DRD2 (Ser311Cys polymorphism) is associated with schizophrenia and worse sustained attention in patients]. Zh Nevrol Psikhiatr Im S S Korsakova. 2009;109(9):67-70. | | 2. Kawanabe Y, Okamoto Y, Hashimoto N, Masaki T: Molecular mechanisms for activation of voltage-independent Ca2+ channels by endothelin-1/endothelin-A receptors. J Cardiovasc Pharmacol. 2004 Nov;44 Suppl 1:S219-23. | | 3. Dupraz P, Oertle S, Meric C, Damay P, Spahr PF: Point mutations in the proximal Cys-His box of Rous sarcoma virus nucleocapsid protein. J Virol. 1990 Oct;64(10):4978-87. |
|
|---|